Genomic analysis of a novel ST11(PR34365) Clostridioides difficile strain isolated from the human fecal of a CDI patient in Guizhou, China
-
Ying Yang
Abstract
Clostridioides difficile is a pathogen that causes pseudomembranous colitis with antibiotic-associated diarrhea. The epidemiology and molecular evolution of C. difficile may differ among different geographic regions, and mining its genomic information can help to understand the epidemiology and molecular evolution of C. difficile and focus on its transmission mode. A C. difficile strain denoted WXL8 was isolated from a human fecal sample from a patient in the intensive care unit, and its physiology and genomic sequence were determined. The total genome size of WXL8 was 4,119,929 bp, and the GC content was 27.97%. The multilocus sequence typing results indicated that WXL8 is strain type 11 (ST11), a genotype widely present in livestock. The WXL8 was located in clade 5 of ST11. The ribotype of WXL8 was a novel ribotype (PR34365). It is the first report of the ST11 (PR34365) strain. Comparative genomic analysis between WXL8 and the other four high-virulence strains (CD630, CDBR81, CDS-0253, and CDR20291) showed differences in gene arrangement, indicating the uniqueness of C. difficile WXL8. In the present study, a novel ribotype (PR34365) ST11 strain of C. difficile was isolated from a patient with diarrhea in Guizhou, China. Our findings suggest that zoonotic CDI should receive more clinical attention.
1 Introduction
Clostridioides difficile is an obligate anaerobic, Gram-positive, spore-forming bacterium found ubiquitously in the environment and the gastrointestinal tracts of humans and animals. It is primarily transmitted through the fecal–oral route [1,2]. C. difficile is the primary pathogen causing antibiotic-associated diarrhea (AAD) in the hospital environment, and it is closely related to the disorder of gut microbiota caused by the use of antibiotics [3,4]. A recent report from the Center for Disease Control and Prevention ranks C. difficile infection (CDI) as the urgent public health threat with 223,900 incidences and 12,800 deaths yearly and the cost of bed days and associated treatment totaling $1 billion. Recently, C. difficile has exceeded multi-drug resistant organisms such as methicillin-resistant Staphylococcus aureus as the urgent-control needed pathogen causing hospital-acquired infection [5,6]. Moreover, the recurrent rate of CDI is estimated to occur at 20–30% [7], which together imposes an extreme burden on the healthcare system worldwide.
A critical development in C. difficile’s molecular evolution is the emergence of binary toxin-producing ribotypes, such as 027 and 176, associated with more severe clinical presentations and higher recurrence rates than non-binary producers. The rapid dissemination and clonal spread of hypervirulent strains like NAP1/RT027 have amplified concerns regarding CDI’s global prevalence, with these strains spreading across Europe, North America, Asia, and Australia [20]. In recent years, there have been substantial changes in C. difficile’s global epidemiology and molecular characteristics. Predominant ribotypes such as 027 and 126 have been replaced by new, highly pathogenic strains with increased virulence factors, resistance profiles, and transmission rates. Ribotype 078 has emerged as a significant cause of CDI outbreaks in several countries, emphasizing the need for continuous monitoring and surveillance [21,22]. The genetic plasticity within C. difficile’s pan-genome has facilitated the exchange of virulence factors and antimicrobial resistance genes among strains, aiding their adaptation in various environments [23]. Transferable C. difficile plasmids harboring antibiotic resistance determinants have been identified in both human and animal populations, highlighting the zoonotic potential of this bacterium. The expanding knowledge on global C. difficile epidemiology and its molecular evolution provides a crucial context for understanding the complex interplay between this bacterium, its human and animal hosts, and the environment [24]. Positioning studies within this broader framework can offer valuable insights into C. difficile’s emergence, transmission, and virulence factors, contributing to improved prevention and control strategies in public health and veterinary medicine.
Genotyping methods are a valuable tool for studying the molecular epidemiology of C. difficile. These methods can help to clarify the genotypes of highly virulent C. difficile strains that are prevalent worldwide [8]. C. difficile sequence type 11 (ST11) is one of the most prevalent and pathogenic sub-lineages of C. difficile [9]. It is primarily associated with livestock (such as chickens, ducks, and pigs) and food contamination [10,11]. In a study of 953 animal fecal samples from China, Zhang et al. isolated 55 C. difficile strains. These strains were classified into three sequence types (STs), with ST11/RT12 being the most common [12]. In another study, Masarikova et al. isolated 44 C. difficile strains from 297 calf fecal samples. Of these strains, 84% were ST11 (RT033) [13]. A study in China also found that ST11 is the predominant ribotype in pigs [9]. These studies suggest that ST11 C. difficile strains are primarily hosted by livestock. We hypothesized that animal-associated strains can be transmitted to humans, which could be an important route of CDI transmission. However, further research is needed to characterize the genotypes and virulence phenotypes of ST11 strains isolated from humans.
With the fast development of sequencing technology, next-generation sequencing (NGS) based on the Illumina NovaSeq platform has enabled us to obtain high-quality bacterial genomic data with more accuracy and speed. The accurate genomic sequence of CDI-causing C. difficile strain has extensively promoted the study of C. difficile gene function and pathogenesis. The well-known model strain of C. difficile, 630, which is virulent and multidrug-resistant, was first sequenced by Sebaihia et al. in 2006 [14]. It was subsequently reannotated by Dupuy [15] and Lawley [16]. Dannheim et al. manually curated the genome sequence of C. difficile strain 630∆erm (DSM 28645), as well as the genome sequence of C. difficile strain 630 (DSM 27543). They completely identified the sequence of the transposon Tn5397 [17]. The genome sequence of C. difficile LCL126, obtained from the Lanzhou Institute of Biological Product, was revealed, along with its encoded proteins and potential toxicological genes [18]. However, extensive research has been carried out on the epidemiology of CDI and molecular characteristics of CDI-causing C. difficile strains. However, there is a general lack of research in the detailed genomic characterization of ST11 C. difficile strains isolated from CDI patients [19].
In the present study, we combined the NGS based on the Illumina NovaSeq platform and third-generation gene sequencing based on the Oxford Nanopore Technologies (ONT) platform to sequence a novel ribotype ST11 strain of C. difficile (ST11, PR34365, denoted WXL8) isolated from a CDI patient in the intensive care unit (ICU) of Affiliated Hospital of Guizhou Medical University. Comparative genomic sequence analyses with model strains were also performed to reveal the differences between typical C. difficile strain and WXL8.
2 Materials and methods
2.1 Isolation of C. difficile WXL8 and retrospective clinical data
Anal swab samples were obtained from patients at the Department of Critical Care Medicine, Affiliated Hospital of Guizhou Medical University in Guiyang, Guizhou, China. Samples were collected using sterile containers and transported under appropriate conditions to prevent contamination and degradation. Swab samples were then soaked in 2 mL of brain heart infusion medium (Solarbio, Lot. 328U032, Beijing, China) for 10 min and -mixed well. A portion of this sample was plated onto Cycloserine-Cefoxitin-Fructose Agar (CCFA), which inhibits the growth of competing flora while promoting the growth of C. difficile, plated and incubated at 37°C within an anaerobic chamber (N2: 95%, H2: 5%) for 48–72 h. Colonies suspected of C. difficile were identified based on their morphology and color on the selective media (light yellow, irregular edges, translucence). They underwent 16S rDNA amplification and sequencing by Sangon Biotech (Shanghai, Co., Ltd, China). The BLAST algorithm [20,21] was employed to classify the isolate taxonomically. Medical records from the Affiliated Hospital of Guizhou Medical University were acquired with patient consent, specifically detailing inpatient treatment and antibiotic usage history.
-
Informed consent: Informed consent has been obtained from all individuals included in this study.
-
Ethical approval: The research related to human use has been complied with all the relevant national regulations, institutional policies and in accordance with the tenets of the Helsinki Declaration, and has been approved by the Ethics committee of Guizhou Medical University (Approval no.: 2021069).
2.2 DNA extraction and genome sequencing
The C. difficile WXL8 strain’s genomic DNA was extracted using the sodium dodecyl sulfate method [22]. The extraction process involved cell lysis, removal of contaminants, and purification of the DNA. The quality and quantity of the extracted DNA were assessed using spectrophotometry and gel electrophoresis to ensure that the DNA was suitable for sequencing. Subsequently, the concentration of the genomic DNA was quantified by applying the Quant-iT PicoGreen dsDNA Assay Kit and the NanoDrop Spectrophotometer (Thermo Scientific, Waltham, USA). Following the successful qualification of the samples, DNA libraries were meticulously constructed. The Whole Genome Shotgun approach was employed to sequence the whole genome of the C. difficile WXL8 strain. Notably, the libraries were subjected to separate sequencing runs utilizing both NGS (Illumina NovaSeq, United States of America, California, NovaSeq 6000) and third-generation single-molecule sequencing technologies (ONT) platform.
2.3 Genome assembly and visualization
The raw sequencing data were processed using bioinformatics tools to assemble the genomes, annotate genes, and identify genetic features relevant to virulence and antibiotic resistance. Raw paired-end sequencing data were initially archived in the FASTQ format (Illumina 1.8+). The assessment of sequencing data quality was carried out using FastQC software [23]. Subsequently, the removal of 3′ adapter contamination was executed through the application of adapter removal [24]. Data underwent filtration based on Kmer frequency (Kmer = 17) using SOAPec (v.2.03) [25], resulting in the extraction of high-quality data from second-generation sequencing, which we refer to as HQdata. Canu software (v1.7.1) [26] was instrumental in assembling the triple sequencing data derived from the ONT platform (Oxford Nanopore Technologies Limited, Oxford, England). Following assembly, all resultant sequences were integrated to construct a comprehensive genome sequence, a process culminating in genome refinement through the utilization of Pilon software [27].
For third-generation sequencing data, assembly into contigs was achieved by employing HGAP [28] and CANU [26]. The HQdata was used to rectify the contigs using Pilon software (v.18) [27]. Finally, the obtained genome sequence, gene prediction, and non-coding RNA prediction information were integrated into a standard GBK (GenBank) format file, and the complete genome of C. difficile WXL8 was assembled using Circos software with predicted condensed proteins for condensed sequences. A circular map of this genome was then drawn using Cgview [29].
2.4 Gene function annotation
Gene prediction was performed employing GeneMarkS software (version 4.32, Georgia Institute of Technology, Atlanta, Georgia, USA), and the resulting amino acid sequences were subjected to a BLAST search against various databases, including gene ontology (GO), Kyoto Encyclopedia of Genes and Genomes (KEGG), clusters of orthologous groups of proteins (COG), carbohydrate-active enzymes database (CAZy), and the pathogen–host interactions database (PHI). Subsequently, a comprehensive filtering and annotation process was implemented.
2.5 Scanning electron microscopy (SEM) and growth curve determination
C. difficile was cultivated in a brain heart infusion-supplemented (BHIS) medium until reaching an optical density at 600 nm (OD600) of 0.6. Subsequently, the bacterial culture was harvested via centrifugation at 3,440 × g for 3 min. The collected bacteria were resuspended in a 2.5% glutaraldehyde fixative solution and left to fix overnight at 4°C. After fixation, the cells underwent a series of washes with phosphate-buffered saline (Gibco, Paisley, Scotland, UK), with each wash being repeated three times. Dehydration of the samples was then accomplished through sequential immersion in ethanol solutions with increasing concentrations: 50, 70, 90, and 100% (v/v), each for 5 min. The dehydrated samples were subsequently subjected to vacuum freeze-drying. Bacteria were delicately isolated from the specimens using a toothpick, affixed to an adhesive substrate, and sputter-coated with a gold film for SEM analysis, conducted with a Hitachi S-3400 SEM instrument (Japan). In parallel, seed C. difficile strain WXL8 strain was cultured to an OD600 of 0.5, then inoculated into BHIS liquid medium with a 1% inoculum and incubated anaerobically at 37°C. Optical density readings at OD600 were recorded at 3 h intervals, enabling the construction of growth curves.
2.6 Multilocus sequence typing (MLST) and capillary gel electrophoresis-based ribotyping
For MLST analysis, we selected housekeeping genes commonly used for C. difficile typing, including adk, atpA, dxr, glyA, recA, sodA, and tpi (Table 1). Specific primers for each selected housekeeping gene were designed based on previously published sequences [30]. PCR amplification was performed using a standard thermal cycler with the following conditions: initial denaturation at 95°C for 5 min, followed by 30 cycles of denaturation at 95°C for 30 s, annealing at 55°C for 30 s, and extension at 72°C for 1 min, concluding with a final extension at 72°C for 5 min. The PCR products were purified using a commercial purification kit (e.g., Qiagen QIAquick PCR Purification Kit) and then sequenced using Sanger sequencing technology. The resulting sequences were analyzed for quality and trimmed to remove low-quality bases. The sequences were compared to C. difficile MLST database (https://pubmlst.org/cdifficile//) to assign alleles and determine the ST of each isolate [9].
MLST
| Strain | adk | atpA | dxr | glyA | recA | sodA | tpi | ST types |
|---|---|---|---|---|---|---|---|---|
| WXL8 | 5 | 8 | 5 | 11 | 9 | 11 | 8 | 11 |
For ribotyping, the 16S–23S rDNA intergenic spacer region of WXL8 was amplified using the primers F1/R1 [31] and F2/R2 [32] (Table 5), which are labeled with 6-carboxyfluorescein (6-FAM) at the 5′ end. The reaction mixture was prepared as follows: 25 μL, including 2× Phanta Flash Master Mix (Dye Plus) (P520, Novozymes, Nanjing, China), 2 μL of each primer, 1 μL of WXL8 genomic DNA (200 ng/μL), and ddH2O to bring the total reaction volume to 25 μL. The PCR cycling conditions were as follows: 30 s of pre-denaturation at 98°C, 10 s of denaturation at 98°C, 5 s of annealing at 57°C, and 1 min of extension at 72°C for 35 cycles, and 1 min of final extension at 72°C. The PCR products were then separated by capillary electrophoresis performed at Sangon Biotech (Shanghai) Co., Ltd. The capillary electrophoresis results (Table 2) were submitted to the Webribo database (http://webribo.ages.at) for C. difficile ribotyping [33].
Capillary gel electrophoresis-based ribotyping
| Fragment a | Fragment b | Fragment c | Fragment d | Fragment e | Fragment f |
|---|---|---|---|---|---|
| 308.02 bp | 324.77 bp | 327.23 bp | 345.33 bp | 370.66 bp | 443.29 bp |
2.7 Construction of phylogenetic tree
Seventy-nine C. difficile strains, including WXL8, were selected for phylogenetic analysis by reference to Dingle et al. [34]. The core genome of 79 strains was identified by clustering analysis using Pirate v1.0.4. Multiple sequence alignment was performed using mafft v7.475, and phylogenetic analysis was conducted using MEGA (MEGA X, Mega Limited, Auckland, New Zealand) to construct a maximum likelihood tree.
2.8 Comparative genomic analysis
In this study, we sought to elucidate the evolutionary trajectory of C. difficile strain WXL8. To achieve this, we retrieved the whole genome sequences of four reference strains, namely C. difficile CD630 (CD630), C. difficile CDS-0253, C. difficile CDR20291, and C. difficile CDBR81, from the National Center for Biotechnology Information. The Mauve software (version 2.4.0, http://darlinglab.org/mauve/mauve.html) was employed to facilitate a comprehensive comparative analysis of the genome of C. difficile WXL8 and these four reference C. difficile strains. Furthermore, Venn plot was generated using R package VennDiagram and Genome Collinearity Chord Diagram was generated using the chord-Diagram function from R package “circlize” (v.0.4.8).
3 Results
3.1 Retrospective clinical data of the host of C. difficile WXL8
A 53-year-old male patient was admitted to the ICU at Guizhou Medical University, presenting with an anterior communicating artery aneurysm and concomitant lung infection. Throughout his hospitalization, the patient underwent an initial therapeutic regimen consisting primarily of glycylcycline and β-lactam antibiotics, administered for 21 days. Notably, after 8 days of antibiotic treatment, the patient experienced the onset of mild diarrhea, prompting the collection of a stool sample via anal swab for further analysis. Subsequent culture of the stool sample on a CCFA plate yielded a distinctive colony exhibiting typical characteristics of C. difficile, characterized by its light-yellow hue, irregular border, and translucency. Subsequent analysis via BLAST (version 2.13.0) revealed a strikingly close genetic affinity between this strain and C. difficile ATCC9689, with an impressive 99.97% sequence identity. Consequently, this isolate was designated as C. difficile WXL8.
3.2 Morphology and growth characteristics of WXL8
The morphological attributes of both the C. difficile CD630 and WXL8 strains were observed using electron microscopy at magnifications of 5,000×, 10,000×, 20,000×, and 30,000×. The observations revealed a noteworthy distinction, with the bacterium of WXL8 displaying greater length when compared to CD630 (average length: 5.65 μm vs 3.58 μm), as illustrated in Figure 1a–i. Additionally, we assessed the growth profiles of WXL8, as depicted in Figure 2a. The growth rate of WXL8 exhibited a rapid increase, reaching its maximum growth concentration (OD600 = 1.76) at 10 h, which was slightly faster than the CD630 strain. Subsequently, at the 12 h, the onset of autolysis of WXL8 became evident, which was earlier than the CD630 strain. Virtual electrophoresis results using genomic DNAs as templates showed that WXL8 had a different ribotype compared to the six typical C. difficile strains: CD630 (RT012), BR81 (RT106), DSM 101085 (RT033), S-0253 (RT002), M120 (RT078), and R20291 (RT027) (Figure 2b).
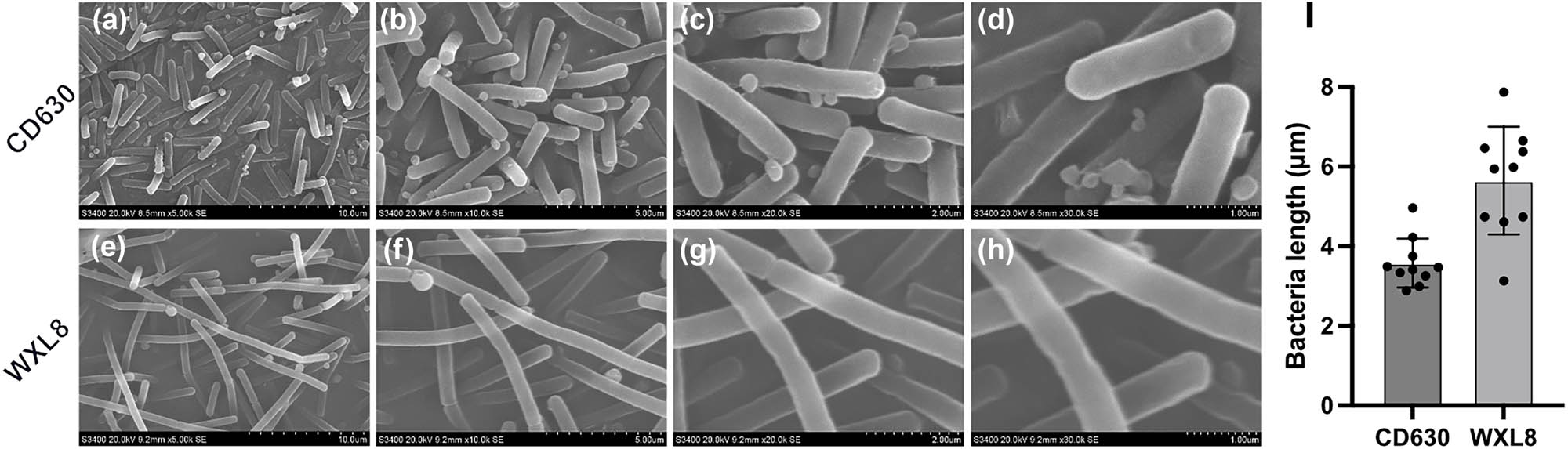

Observation of cell morphology of WXL8 via SEM. (a)–(d) Cell morphology of C. difficile CD630 at magnifications of 5,000 ×, 10,000 ×, 20,000 ×, and 30,000 ×, respectively. (e)–(h) Cell morphology of WXL8 at the same magnifications. Additionally, (i) the average length of WXL8 is compared to C. difficile CD630.

Growth curve and ribotyping analysis of WXL8. (a) The x-axis illustrates the culture time, while the y-axis represents the optical density (OD600) values measured at 600 nm. (b) A comparison between the electronic gel electrophoresis pattern of WXL8 and the well-known ST11 subribotypes is presented. (b) Virtual electrophoresis profiles of PCR ribotyping of seven C. difficile strains. M is a 1 kb Plus DNA Ladder (New England Biolabs). Lane 1 is CD630 (RT012), lane 2 is BR81 (RT106), lane 3 is DSM 101085 (RT033), lane 4 is S-0253 (RT002), lane 5 is M120 (RT078), lane 6 is R20291 (RT027), and lane 7 is WXL8.
3.3 Genome characteristics of C. difficile WXL8
High-throughput sequencing data provided comprehensive insights into the genomic characteristics of C. difficile WXL8. The circular genome of C. difficile WXL8 was determined to encompass a total size of 4,119,929 base pairs (bp) with a GC content of 29.06% (GenBank Accession Number: CP137339). Notably, our examination did not reveal the presence of any plasmids within its genome, as illustrated in Figure 3. The WXL8 genome contains a total of 3,752 predicted protein-coding genes, with an average length of 906.28 bp (accounting for 82.53% of the total length) (Figure 3a). To visualize the circular structure of the WXL8 genome, we utilized the cgview tool, and the graphical representation is presented in Figure 3b. The genomic content also encompassed a diverse repertoire of 271 RNA copies, including 35 rRNA copies (11 copies of 5S rRNA, 12 copies of 16S rRNA, and 23S rRNA), 89 tRNA genes, and 147 non-coding RNAs.
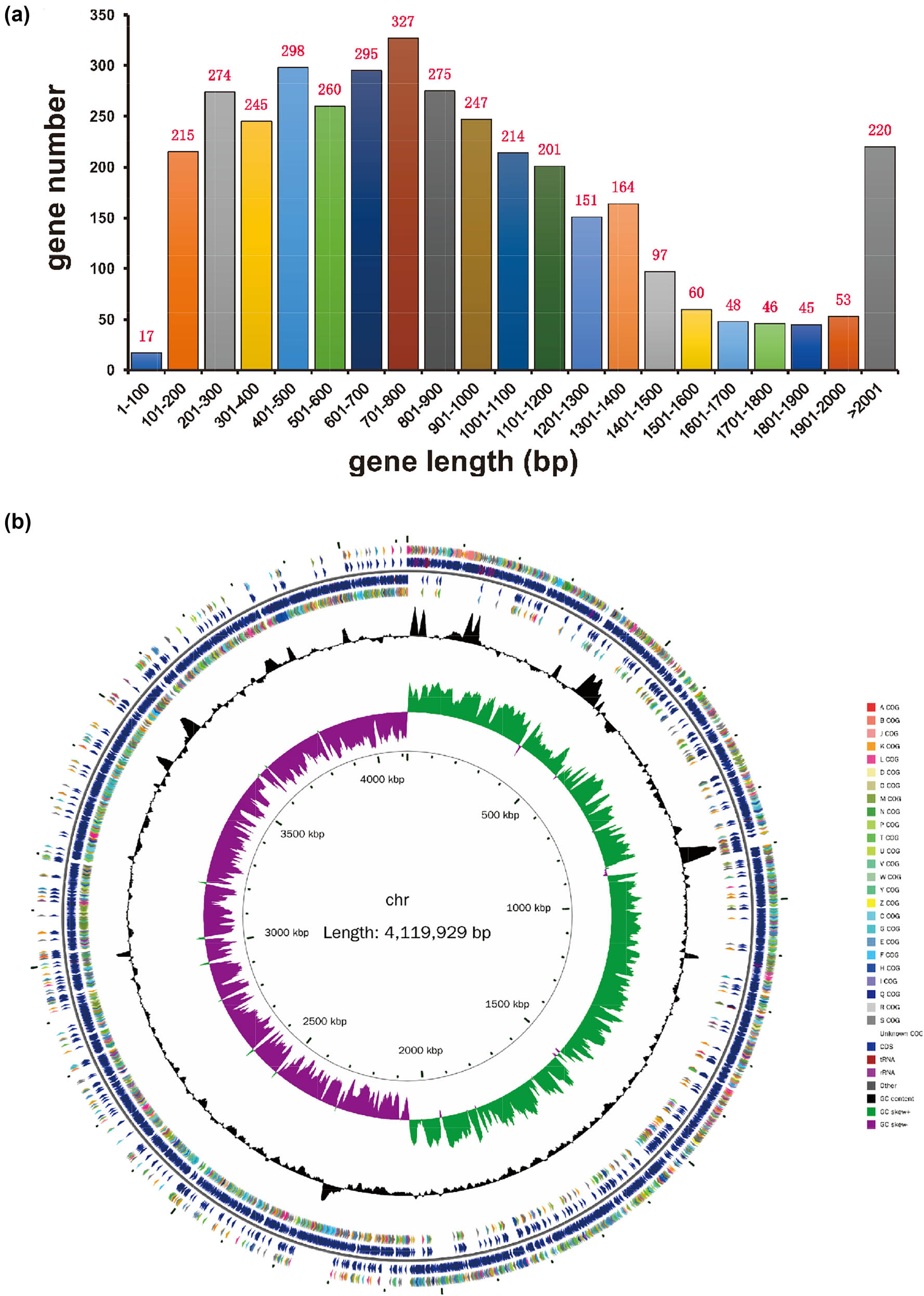
Gene length statistics and genome circle map of WXL8. (a) The x-axis represents gene length, while the y-axis shows the corresponding gene count. (b) In the circular representation, the innermost circle denotes the scale, followed by the depiction of GC Skew, GC content, and the COG of each CDS from the fourth to seventh circles. The fifth and sixth circles provide the positional information of CDS, tRNA, and rRNA within the genome.
Additionally, we identified a total of 178 repetitive sequences, comprising 27 long interspersed nuclear elements, 88 long terminal repeats, 39 DNA transposons, 2 satellite RNAs, and 8 unclassified interspersed repeats. Our investigation also detected 16 prophages within the genome, collectively spanning a total length of 407,296 bp, with an average prophage length of 25,456 bp. Furthermore, we predicted the presence of 15 clustered regularly interspaced short palindromic repeats (CRISPRs) arrays, each characterized by an average length of 777.4 bp. Lastly, our analysis revealed the prediction of 33 genomic islands (GIs), with a cumulative length of 440,363 bp and an average length of 16,310 bp. Detailed information regarding the spatial distribution of these gene islands can be found in Figure S1.
3.4 MLST typing and ribotyping
The genome sequence of WXL8 was uploaded to the MLST database. As shown in Table 1, the results indicated that the MLST type of WXL8 was ST11, which is common in livestock but relatively rare in humans [35]. The ribotype identification results, as shown in Table 2, indicated that the WXL8 strain was defined as a novel ribotype, PR34365, by the Webribo database.
3.5 Genome functional analysis of C. difficile WXL8
To elucidate the functional aspects of the C. difficile WXL8 genome, a comprehensive investigation encompassing gene annotation, effector profiling, and virulence analysis was conducted. GO annotation yielded a substantial dataset, revealing a total of 16,603 functionally classified terms. Within this dataset, three primary ontology categories were identified: biological process (2,543 terms), cellular component (825 terms), and molecular function (2,540 terms), as depicted in Figure 4. These categorizations collectively constitute a comprehensive framework for comprehending the multifaceted gene functions within the genome of C. difficile WXL8.

GO functional annotation across the entire genome of WXL8. The x-axis denotes the number of genes in the sample annotation, while the y-axis represents the GO functional classification.
The gene function exploration of C. difficile WXL8 was further enriched through KEGG annotation, which revealed a comprehensive dataset comprising 3,311 genes. These genes were thoughtfully categorized into eight distinct pathways, delineated as Brite Hierarchies, Cellular Processes, Environmental Information Processing, Genetic Information Processing, Human Diseases, Metabolism, Not Included in Pathway or Brite, and Organismal Systems (Figure 5). Among these pathways, notable prominence was observed in various categories. In the Brite Hierarchies class, 498 unigenes were associated with protein families predominantly influencing genetic information processing. In the Cellular Processes class, 66 unigenes played pivotal roles in cellular community-prokaryotes, while the Environmental Information Processing class featured 148 unigenes notably associated with membrane transport. Within the Genetic Information Processing class, 82 unigenes held significance in the domain of translation, further enriching our understanding of this fundamental process. The Human Diseases class unveiled 35 unigenes contributing to drug resistance, particularly in the context of antimicrobial resistance. Metabolism, a critical aspect of cellular function, was notably represented by 374 unigenes in the metabolism class, with a predominant focus on carbohydrate metabolism. In the Not Included in Pathway or Brite class, 119 unigenes were identified, primarily associated with unclassified metabolism. Lastly, in the Organismal Systems class, 13 unigenes were intricately linked to the endocrine system, further expanding our comprehension of the genomic contributions to this intricate biological system. This comprehensive categorization provides a valuable framework for deciphering the multifaceted functional aspects of genes within the C. difficile WXL8 genome.

Distribution of genes based on the KEGG classification in WXL8. The numbers on the x-axis indicate the count of genes in the annotation, while the y-axis represents the distinct functional categories.
In the COG database, we identified 3,374 genes and classified them into 24 distinct categories. The most abundant category was Transcription, which houses 333 genes, followed closely by Amino Acid Transport and Metabolism and Carbohydrate Transport and Metabolism, which included 265 and 254 genes, respectively (Figure 6a). Additionally, 69 genes were associated with defense mechanisms, while the functions of 928 genes remained uncharted. In the CAZy database, we annotated 102 genes, comprising 45 glycoside hydrolases, 25 glycosyl transferases, 21 carbohydrate esterases, 9 auxiliary activities, and 2 carbohydrate-binding modules (Figure 7). Furthermore, we identified 77 signal peptides, 961 transmembrane structures, and 94 secreted proteins. These secreted proteins are pivotal in the context of infection, where they play a crucial role in the intricate pathogen–host interactions that underlie CDI.

Annotated clusters based on the COG in WXL8 and the mutated genes in the PHI. (a) Functional classification is represented along the x-axis, while the count of genes is shown on the y-axis. (b) Function classes are indicated along the x-axis, and the number of mutated genes is represented on the y-axis.

CAZy functional classification chart for WXL8. The x-axis denotes the number of genes in the sample annotation, while the y-axis represents the different CAZy functional categories.
Analysis in the Virulence Factors of Pathogenic Bacteria (VFDB) database revealed that the C. difficile WXL8 strain carries nine toxin genes, including Toxin A, Toxin B, UDP-N-acetylglucosamine-2-epimerase, CdtA, CdtB, lota-toxin components Ia and Ib, along with two copies of ATP-dependent Clp protease proteolytic subunit (Table 3). Furthermore, the PHI examination indicated that 495 genes could potentially reduce toxicity, while 94 genes had the opposite effect (Figure 6b). In the Comprehensive Antibiotic Resistance Database (CARD) analysis, a total of 35 genes were annotated, encompassing 22 antibiotic resistance genes and 17 antibiotic target genes (Table 4). These findings shed light on the diverse genomic attributes of C. difficile WXL8, which contribute to its functional repertoire and interactions with its host environment.
VFDB result statistics
| VFDB ID | ORF name | VFDB name |
|---|---|---|
| VFG002288(gb|YP_001087135) | chr_689 | VF0377 |
| VFG002287(gb|YP_001087137) | chr_692 | VF0376 |
| VFG001373(gb|NP_344890) | chr_1022 | VF0144 |
| VFG002298(gb|AAF81760) | chr_2636 | VF0385 |
| VFG002292(gb|CAA51959) | chr_2636 | VF0381 |
| VFG002299(gb|AAF81761) | chr_2637 | VF0385 |
| VFG002293(gb|CAA51960) | chr_2637 | VF0381 |
| VFG000077(gb|NP_465991) | chr_3370 | VF0074 |
| VFG000077(gb|NP_465991) | chr_3395 | VF0074 |
Statistics of antibiotic resistance analysis results
| Seq ID | Property | Number of genes | Percentage |
|---|---|---|---|
| chr | Antibiotic resistance | 22 | 0.586 |
| Antibiotic target | 17 | 0.453 | |
| Antibiotic biosynthesis | 0 | 0.000 | |
| Total genes | 35 | 0.933 |
3.6 Phylogenetic analysis
To explore the phylogenetic status of WXL8, we constructed a phylogenetic tree that incorporated WXL8 using the neighbor-joining method. The resulting phylogenetic tree showcased representatives of C. difficile species from the six recognized evolutionary branches and illuminated relationships between virulence-producing and non-virulence-producing isolates. In Figure 8, WXL8, along with four ST11 strains as well as ST164, ST167, and ST168, cluster in clade 5, holding a unique position in the evolutionary tree. Notably, all ST11 strains within this branch exhibit the tcdA + and tcdB + phenotype.

Phylogenetic analysis. The evolutionary relationships of WXL8 and typical C. difficile strains were depicted by a phylogenetic tree using the Neighbor-Joining method. Green triangles denote virulence-producing strains, while non-virulence-producing strains are marked with red circles. Different clades are depicted in different colors. The scale of the tree is 0.01.
3.7 Comparative genomic analyses
Through collinear gene comparisons, we meticulously examined the consistency and variability among the whole genomes of C. difficile CD630 [14], CDBR81 [36], CDS-0253 [37], CDR20291 [38], and WXL8 strains, offering insights into their shared genetic ancestry. The comprehensive gene covariance analysis, as illustrated in Figure 9, enabled a closer inspection of WXL8 in comparison to the other four C. difficile strains. In terms of gene structure alignment, WXL8 exhibited a higher similarity to the reference genomes C. difficile CD630 and C. difficile CDR20291. Specifically, the gene structure arrangement in WXL8 closely resembled that of CDR20291, although notable distinctions were observed compared to C. difficile CDBR81.

Multi-gene sequence comparison of strains WXL8, CD630, CDS-0253, CDR20291, and CDBR81. Mauve alignment of four representative complete genomes within each of the four species of C. difficile: CD630, CDBR81, WXL8, CDS-0253, and CDR20291. Blocks of the same color correspond to locally collinear blocks. Each sequence of identically colored blocks represents a collinear set of matching regions. One connecting line is drawn per collinear block.
The pan-genome encompasses all genes within a species. It comprises three main components: core genes (present in all strains), dispensable genes (not essential but present in some strains), and strain-specific unique genes [39]. We examined the genomic distribution of five C. difficile strains – 630, R20291, S-0253, BR81, and WXL8 – using a Venn diagram (Figure S2a) to determine the number of core and unique genes across samples. The results revealed that WXL8 contains 4,372 genes, with 3,096 shared with the strains 630, R20291, S-0253, and BR81. Notably, C. difficile strain WXL8 possesses the highest number of unique genes (253) among the five examined strains. The R20291 strain has 155 unique genes, while strain 630 has 123, strain S-0253 contains 74, and strain BR81 exhibits the fewest, with only 53 unique genes. This indicates that WXL8 displays a distinct genomic profile compared to other strains.
Synteny refers to the conservation of gene order on chromosomes across different species. Generally, more distantly related species exhibit less gene synteny, thus serving as a metric for evolutionary distance. We conducted a synteny analysis between the genomes of C. difficile 630 and WXL8 to explore their genomic relationships. The results are depicted in Figure S2b, where colored lines illustrate connections between the two genomes; blue represents C. difficile 630, while orange denotes C. difficile WXL8. This result representation effectively highlights the corresponding genomic regions between WXL8 and C. difficile strain 630. Overall, WXL8 exhibits similarity to the reference genomes of C. difficile CD630 (Figure S2b).
A partial deletion was observed within the 0.4–0.6 Mb range of WXL8 compared to the other four genomes analyzed. In contrast to CD630, WXL8 exhibited a single reverse fragment insertion spanning 2.2–2.4 Mb and a double fragment insertion between 3.6 and 3.8 Mb. Interestingly, one fragment displayed a forward orientation while the other was reversed, contributing to genomic diversity. Additionally, an inversion event spanning 3.4–3.6 Mb was identified in WXL8 compared to CD630. Further investigation revealed inversions at the 2.2–2.4 Mb locus relative to the CDBR81 genome and fragment inversions within the 3.6–3.8 Mb region compared to the C. difficile CDR20291 genome. These comparative genomic analyses highlight the subtle variations and structural rearrangements that distinguish WXL8 from its closely related counterparts.
4 Discussion
C. difficile toxin is a major pathogenic factor associated with the occurrence of CDI in hospitals [40]. However, C. difficile can be isolated not only in human feces but also in a variety of animals (pigs, cattle, chickens, ducks, etc.), and with the emergence of highly virulent strains of C. difficile, CDI has now become a significant public health problem internationally [41]. In the present study, a strain of C. difficile, named WXL8 (PR34365), isolated from the feces of patients in the ICU ward of the Affiliated Hospital of Guizhou Medical University, was observed morphologically by SEM and its growth curve was measured. The whole genome sequence of WXL8 (PR34365) was obtained by high-throughput sequencing. The MLST typing and capillary gel electrophoresis-based ribotyping categorized it as ST11 genotype and PR34365 ribotype. A phylogenetic tree was constructed after extracting core genes, and comparative genomic analysis was performed with other C. difficile strains. The VFDB annotation results showed that WXL8 contained TcdA, TcdB, and the binary toxin CdtA/CdtB. In addition, 94 genes showed increased virulence in the PHI annotation results and 22 antibiotic resistance genes in the CARD annotation results. To our limited knowledge, the WXL8 is the first ST11(PR34365) C. difficile strain isolated from a CDI patient.
C. difficile sequence type 11 (ST11) is recognized for its global prevalence in infecting and colonizing livestock. In 2005, Goorhuis et al. first reported the isolation of human ST11 C. difficile [42]. It encompasses a diverse array of ribotypes, several of which are implicated in human diseases. This pattern indicates that CDIs could potentially be a zoonosis [43]. ST11 interests researchers as one of the world’s most widespread and threatening classes of genotypes, especially in Asia [44]. C. difficile RT078, a notable sub-lineage within ST11, has established significant reservoirs in livestock globally [45]. It has also been detected in various retail meat products across North America and Europe [46]. It is a leading cause of both hospital-acquired and community-acquired CDI in these regions [42,47,48]. However, previous studies have shown that ST11 CDI cases are rarely detected in China [49]. In 2021, Zhang et al. isolated 99 strains of C. difficile from diarrhea patients under 5 years of age, with the main genotypes being ST3, ST54, and ST35, and only one strain was ST11/RT078 (1%) [50]. The same research group isolated 116 strains of C. difficile from elderly fecal samples in 2018, of which 14 were ST11 (12%). In the study by Wang et al., 46 strains of C. difficile were isolated, with the primary genotype being ST54, and no ST11 strain was isolated [51]. The transmission route and epidemiological characteristics of the ST11 ribotype in CDI patients in China are still unclear [52]. This study suggests that ST11, which is common in livestock, can infect humans, confirming its zoonotic nature. Therefore, people who come into contact with livestock should be aware of the risk of CDI.
WXL8 was typed by MLST and capillary gel electrophoresis-based ribotyping and was known to belong to ST11(PR34365). The core genome of WXL8 was extracted, and the phylogenetic tree was constructed with 78 other C. difficile individuals. The results show that WXL8 is located on clade 5, a unique branch and that strains on this branch are mostly ST11 genotypes and present toxin genes. In China, Clade 1 is the dominant group of clinical isolates, while Clade 5 is relatively rare [53,54]. The virtual and capillary gel electrophoresis results showed that WXL8 had a unique ribotype. To be cautious in identifying the new ribotype, we repeated the analysis using two sets of primers (F1/R1, F2/R2, Table 5) labeled with 6-FAM. The results of both sets of primers were consistent with the PR34365 ribotype. This ribotype was first reported in ST11, and its clinical epidemiology needs to be further studied in future work.
Strains and primers
| Strain/primer | Relevant characteristic | Source |
|---|---|---|
| C. diffcile 630(AM180355.1) | 4.29 Mb, 3,981 genes, 29.0% G + C content | Switzerland |
| C. difficile BR81(CP019870.1) | 4.12 Mb, 3,747 genes, 28.5% G + C content | Korea |
| C. difficile s-0253(CP076401.1) | 4.08 Mb, 3,732 genes, 28.5% G + C content | Australia |
| C. difficile R20291(CP029423.1) | 4.20 Mb, 3,843 genes, 29.0% G + C content | United Kingdom |
| C. difficile WXL8(CP137339) | 4.11 Mb, 3,752 genes, 29.0% G + C content | China |
| F1 | 5′-GTGCGGCTGGATCACCTCCT-3′ | [31] |
| R1 | 5′-CCCTGCACCCTTAATAACTTGACC-3′ | |
| F2 | 5′-GCTGGATCACCTCCTTTCTAAG-3′ | [32] |
| R2 | 5′-TGACCAGTTAAAAAGGTTTGATAGATT-3′ |
The comparative genomic analysis indicated that WXL8 has a unique gene arrangement compared to other strains such as CD630, CDS-0253, CDR20291, and CDBR81. The genomic analysis revealed that WXL8 harbors nine toxin genes (Table 3), including Toxin A (TcdA), Toxin B (TcdB), and the binary toxin genes (CdtA and CdtB). These toxins are critical for the pathogenicity of C. difficile, as they disrupt the host’s intestinal epithelial cells, leading to inflammation and diarrhea. The presence of these toxins suggests that WXL8 has the potential to cause severe disease, similar to other hypervirulent strains like RT027 and RT078. For instance, the presence of both tcdA and tcdB suggests a potential for severe disease manifestation, while the binary toxin (cdtA/cdtB) is associated with increased virulence and has been implicated in more severe clinical outcomes [55]. The CARD analysis revealed that WXL8 carries 22 antibiotic resistance genes and 17 antibiotic target genes, suggesting that WXL8 has a robust arsenal of resistance mechanisms. These genes could confer resistance to multiple antibiotics, including those commonly used to treat CDIs, such as metronidazole and vancomycin. The 15 CRISPR arrays identified in the WXL8 genome may play a role in antibiotic resistance by providing a defense mechanism against foreign genetic elements, such as plasmids carrying antibiotic resistance genes. This could allow WXL8 to maintain its resistance profile and potentially acquire new resistance genes through horizontal gene transfer. The presence of these resistance genes may explain the high recurrence rate of CDI in patients, as antibiotic pressure could be selected for resistant strains like WXL8.
Concern host interaction, first, the genome of WXL8 encodes 94 secreted proteins, which are crucial for host–pathogen interactions. These proteins may include enzymes that degrade host tissues, immune modulators that evade the host’s immune response, or adhesins that facilitate colonization of the intestinal mucosa. The presence of these secreted proteins suggests that WXL8 has evolved mechanisms to interact with and manipulate the host environment, enhancing its ability to cause infection. Second, the KEGG annotation revealed that WXL8 has a significant number of genes involved in carbohydrate metabolism, which could be crucial for its survival in the gut environment. The ability to efficiently metabolize various carbohydrates may give WXL8 a competitive advantage in the gut microbiome, allowing it to outcompete other bacteria and establish infection. Moreover, the identification of 33 GIs in WXL8, some of which may carry virulence-related genes, could contribute to its pathogenicity. These GIs might encode additional virulence factors or regulatory elements that enhance the strain’s ability to colonize and infect the host.
Identifying ribotype ST11(PR34365) may indicate a potential reservoir in animal populations, possibly a source of human infection. Understanding the host range of this ribotype is crucial for assessing its zoonotic potential. Investigating which animal species carry this ribotype can help identify potential reservoirs; for instance, if ST11(PR34365) is found in livestock, wildlife, or companion animals, it raises concerns about human transmission routes, especially in settings where humans and animals interact closely. The study of environmental and ecological factors that facilitate the transmission of ST11(PR34365) from animals to humans is essential, including examining food production systems, handling practices, and environmental contamination. The emergence of a novel ribotype with zoonotic potential can also have significant public health implications, necessitating enhanced surveillance in both animal and human populations. Monitoring for this ribotype in clinical settings can help in the early detection of zoonotic infections. Understanding the transmission dynamics of ST11(PR34365) can inform infection control measures in healthcare settings, particularly in hospitals where patients may be at higher risk of infection from zoonotic pathogens. Further conducting epidemiological studies to track the incidence of infections associated with ST11(PR34365) can help establish its role in zoonotic transmission while investigating the ecological niches and environmental conditions that favor the persistence and spread of ST11(PR34365) and can inform strategies to mitigate its impact. Continued research in this area is essential to safeguard public health and enhance our understanding of the complex interactions among humans, animals, and the environment.
Guizhou is a predominantly rural province with a significant agricultural sector, including livestock farming. The close interaction between humans and animals in this region increases the risk of zoonotic transmission of C. difficile. Public health interventions should be tailored to the specific needs of rural communities, including education on the risks of zoonotic diseases and the promotion of safe farming practices. The healthcare infrastructure in Guizhou, particularly in rural areas, may be less equipped to handle outbreaks of CDI. Strengthening healthcare capacity, including access to diagnostic tools and effective treatments, is essential to mitigate the impact of zoonotic C. difficile transmission. In healthcare settings, particularly in rural hospitals in Guizhou, infection control measures should be strengthened to prevent the spread of C. difficile. This includes proper hand hygiene, environmental cleaning, and the judicious use of antibiotics to reduce the risk of CDI. Public health campaigns should also educate farmers and the general public of Guizhou, China, about the risks of zoonotic transmission and the importance of good hygiene practices.
The findings of this study on the novel ST11(PR34365) C. difficile strain WXL8 both align with and differ from previous research on ST11 and other ribotypes. While the association with livestock, virulence factors, and antibiotic resistance are consistent with previous studies, identifying a novel ribotype and isolating a human patient in China provide new insights into the genetic diversity and zoonotic potential of the ST11 lineage. These findings underscore the importance of continued surveillance and research to understand the epidemiology, virulence, and resistance profiles of C. difficile strains, particularly in regions with close human–animal interaction. This study enriched the genomic data on C. difficile and laid the foundation for future exploration of the molecular epidemiology of C. difficile in Guizhou, China.
-
Funding information: This work was supported by the National Natural Science Foundation of China (32170134), the Outstanding Young Talent Program of Guizhou Medical University ([2022]101), the Innovation and Entrepreneurship Program for overseas talents (202208), and Guizhou Province High-level Innovative Talent Program Qiankehe Platform Talent-GCC[2023]080.
-
Author contributions: Ying Yang, Luhong Shu, Ping Ling, and Junyi Yang performed the experiments, analyzed the data, and created the figures. Ruirui Shao, Yumei Cheng, Shanshan Luo, and Xinglang Wei performed experiments and provided technical support for imaging analyses. Zhizhong Guan, Zhenghong Chen, Jian Liao, Xiaolan Qi, and Guzhen Cui critically revised the manuscript for important intellectual content. Wei Hong contributed to the study’s design, provided materials, and revised the manuscript. All authors approved the final manuscript.
-
Conflict of interest: Authors state no conflict of interest.
-
Data availability statement: The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
[1] Suetens C, Latour K, Kärki T, Ricchizzi E, Kinross P, Moro ML, et al. Prevalence of healthcare-associated infections, estimated incidence and composite antimicrobial resistance index in acute care hospitals and long-term care facilities: results from two European point prevalence surveys, 2016 to 2017. Euro Surveill Bull Eur Sur Mal Transm Eur Commun Dis Bull. 2018 Nov;23(46):1800516.10.2807/1560-7917.ES.2018.23.46.1800516Search in Google Scholar PubMed PubMed Central
[2] Yassin SF, Young-Fadok TM, Zein NN, Pardi DS. Clostridium difficile-associated diarrhea and colitis. Mayo Clin Proc. 2001 Jul;76(7):725–30.10.4065/76.7.725Search in Google Scholar PubMed
[3] Smits WK, Lyras D, Lacy DB, Wilcox MH, Kuijper EJ. Clostridium difficile infection. Nat Rev Dis Primer. 2016 Apr;2:16020.10.1038/nrdp.2016.20Search in Google Scholar PubMed PubMed Central
[4] Slimings C, Riley TV. Antibiotics and hospital-acquired Clostridium difficile infection: update of systematic review and meta-analysis. J Antimicrob Chemother. 2014 Apr;69(4):881–91.10.1093/jac/dkt477Search in Google Scholar PubMed
[5] Lessa FC, Mu Y, Bamberg WM, Beldavs ZG, Dumyati GK, Dunn JR, et al. Burden of Clostridium difficile infection in the United States. N Engl J Med. 2015 Feb;372(9):825–34.10.1056/NEJMoa1408913Search in Google Scholar PubMed PubMed Central
[6] Evans CT, Fitzpatrick M, Ramanathan S, Kralovic SM, Burns SP, Goldstein B, et al. Healthcare facility-onset, healthcare facility-associated Clostridioides difficile infection in Veterans with spinal cord injury and disorder. J Spinal Cord Med. 2020 Sep;43(5):642–52.10.1080/10790268.2019.1672953Search in Google Scholar PubMed PubMed Central
[7] Bakken JS, Polgreen PM, Beekmann SE, Riedo FX, Streit JA. Treatment approaches including fecal microbiota transplantation for recurrent Clostridium difficile infection (RCDI) among infectious disease physicians. Anaerobe. 2013 Dec;24:20–4.10.1016/j.anaerobe.2013.08.007Search in Google Scholar PubMed
[8] Clabots CR, Johnson S, Bettin KM, Mathie PA, Mulligan ME, Schaberg DR, et al. Development of a rapid and efficient restriction endonuclease analysis typing system for Clostridium difficile and correlation with other typing systems. J Clin Microbiol. 1993 Jul;31(7):1870–5.10.1128/jcm.31.7.1870-1875.1993Search in Google Scholar PubMed PubMed Central
[9] Zhang LJ, Yang L, Gu XX, Chen PX, Fu JL, Jiang HX. The first isolation of Clostridium difficile RT078/ST11 from pigs in China. PLoS One. 2019;14(2):e0212965.10.1371/journal.pone.0212965Search in Google Scholar PubMed PubMed Central
[10] Stabler RA, Dawson LF, Valiente E, Cairns MD, Martin MJ, Donahue EH, et al. Macro and micro diversity of Clostridium difficile isolates from diverse sources and geographical locations. Ahmed N, editor. PLoS One. 2012 Mar;7(3):e31559.10.1371/journal.pone.0031559Search in Google Scholar PubMed PubMed Central
[11] Knight DR, Riley TV. Genomic delineation of zoonotic origins of Clostridium difficile. Front Public Health. 2019 Jun;7:164.10.3389/fpubh.2019.00164Search in Google Scholar PubMed PubMed Central
[12] Zhang WZ, Li WG, Liu YQ, Gu WP, Zhang Q, Li H, et al. The molecular characters and antibiotic resistance of Clostridioides difficile from economic animals in China. BMC Microbiol. 2020;20(1):70.10.1186/s12866-020-01757-zSearch in Google Scholar PubMed PubMed Central
[13] Masarikova M, Simkova I, Plesko M, Eretova V, Krutova M, Cizek A. The colonisation of calves in Czech large-scale dairy farms by clonally-related Clostridioides difficile of the sequence type 11 represented by ribotypes 033 and 126. Microorganisms. 2020 Jun;8(6):901.10.3390/microorganisms8060901Search in Google Scholar PubMed PubMed Central
[14] Sebaihia M, Wren BW, Mullany P, Fairweather NF, Minton N, Stabler R, et al. The multidrug-resistant human pathogen Clostridium difficile has a highly mobile, mosaic genome. Nat Genet. 2006 Jul;38(7):779–86.10.1038/ng1830Search in Google Scholar PubMed
[15] Monot M, Boursaux-Eude C, Thibonnier M, Vallenet D, Moszer I, Medigue C, et al. Reannotation of the genome sequence of Clostridium difficile strain 630. J Med Microbiol. 2011 Feb;60:1193–9.10.1099/jmm.0.030452-0Search in Google Scholar PubMed
[16] Pettit LJ, Browne HP, Yu L, Smits WK, Fagan RP, Barquist L, et al. Functional genomics reveals that Clostridium difficile Spo0A coordinates sporulation, virulence and metabolism. BMC Genomics. 2014;15:160.10.1186/1471-2164-15-160Search in Google Scholar PubMed PubMed Central
[17] Dannheim H, Riedel T, Neumann-Schaal M, Bunk B, Schober I, Spröer C, et al. Manual curation and reannotation of the genomes of Clostridium difficile 630Δerm and C. difficile 630. J Med Microbiol. 2017 Mar;66(3):286–93.10.1099/jmm.0.000427Search in Google Scholar PubMed
[18] Wang J, Yang C, Zhang C, Mao X, Lizhe A. Complete genome sequence of the Clostridium difficile LCL126. Bioengineered. 2021 Dec;12(1):745–54.10.1080/21655979.2021.1894798Search in Google Scholar PubMed PubMed Central
[19] Bloomfield LE, Riley TV. Epidemiology and risk factors for community-associated Clostridium difficile infection: a narrative review. Infect Dis Ther. 2016 Sep;5(3):231–51.10.1007/s40121-016-0117-ySearch in Google Scholar PubMed PubMed Central
[20] Hong W, Cheng Y, Rao F, Yang J, Cui G, Chen Z, et al. Co-infection of Clostridioides (Clostridium) difficile GMU1 and Bacillus cereus GMU2 in one patient in Guizhou, China. Anaerobe. 2018 Dec;54:159–63.10.1016/j.anaerobe.2018.08.004Search in Google Scholar PubMed
[21] Hong W, Yang J, Cheng Y, Huang X, Rao F, Zhang T, et al. Bacteria co-colonizing with Clostridioides Difficile in two asymptomatic patients. Open Life Sci. 2019 Jan;14:628–37.10.1515/biol-2019-0071Search in Google Scholar PubMed PubMed Central
[22] Wilson K. Preparation of genomic DNA from bacteria. Curr Protoc Mol Biol. 2001 Nov;Chapter 11:144–51.Search in Google Scholar
[23] Patel RK, Jain M. NGS QC toolkit: a toolkit for quality control of next generation sequencing data. PLoS One. 2012;7(2):e30619.10.1371/journal.pone.0030619Search in Google Scholar PubMed PubMed Central
[24] Schubert M, Lindgreen S, Orlando L. AdapterRemoval v2: rapid adapter trimming, identification, and read merging. BMC Res Notes. 2016 Dec;9(1):88.10.1186/s13104-016-1900-2Search in Google Scholar PubMed PubMed Central
[25] Luo R, Liu B, Xie Y, Li Z, Huang W, Yuan J, et al. SOAPdenovo2: An empirically improved memory-efficient short-read de novo assembler. GigaScience. 2012 Dec;1(1):18.10.1186/2047-217X-1-18Search in Google Scholar PubMed PubMed Central
[26] Koren S, Walenz BP, Berlin K, Miller JR, Bergman NH, Phillippy AM. Canu: scalable and accurate long-read assembly via adaptive k-mer weighting and repeat separation. Genome Res. 2017 May;27(5):722–36.10.1101/gr.215087.116Search in Google Scholar PubMed PubMed Central
[27] Walker BJ, Abeel T, Shea T, Priest M, Abouelliel A, Sakthikumar S, et al. Pilon: an integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS One. 2014;9(11):e112963.10.1371/journal.pone.0112963Search in Google Scholar PubMed PubMed Central
[28] Phased diploid genome assembly with single-molecule real-time sequencing - PubMed [Internet]. [cited 2023 Mar 13]. https://pubmed.ncbi.nlm.nih.gov/27749838/.Search in Google Scholar
[29] Stothard P, Wishart DS. Circular genome visualization and exploration using CGView. Bioinformatics. 2005 Feb;21(4):537–9.10.1093/bioinformatics/bti054Search in Google Scholar PubMed
[30] Griffiths D, Fawley W, Kachrimanidou M, Bowden R, Crook DW, Fung R, et al. Multilocus sequence typing of Clostridium difficile. J Clin Microbiol. 2010 Mar;48(3):770–8.10.1128/JCM.01796-09Search in Google Scholar PubMed PubMed Central
[31] Bidet P, Barbut F, Lalande V, Burghoffer B, Petit JC. Development of a new PCR-ribotyping method for Clostridium difficile based on ribosomal RNA gene sequencing. FEMS Microbiol Lett. 1999 Jun;175(2):261–6.10.1111/j.1574-6968.1999.tb13629.xSearch in Google Scholar PubMed
[32] Janežič S, Štrumbelj I, Rupnik M. Use of modified PCR ribotyping for direct detection of Clostridium difficile ribotypes in stool samples. J Clin Microbiol. 2011 Aug;49(8):3024–5.10.1128/JCM.01013-11Search in Google Scholar PubMed PubMed Central
[33] Fawley WN, Knetsch CW, MacCannell DR, Harmanus C, Du T, Mulvey MR, et al. Development and validation of an internationally-standardized, high-resolution capillary gel-based electrophoresis PCR-ribotyping protocol for Clostridium difficile. Friedrich A, editor. PLoS One. 2015 Feb;10(2):e0118150.10.1371/journal.pone.0118150Search in Google Scholar PubMed PubMed Central
[34] Dingle KE, Elliott B, Robinson E, Griffiths D, Eyre DW, Stoesser N, et al. Evolutionary history of the Clostridium difficile pathogenicity locus. Genome Biol Evol. 2014 Jan;6(1):36–52.10.1093/gbe/evt204Search in Google Scholar PubMed PubMed Central
[35] Dingle KE, Didelot X, Ansari MA, Eyre DW, Vaughan A, Griffiths D, et al. Recombinational switching of the Clostridium difficile S-layer and a novel glycosylation gene cluster revealed by large-scale whole-genome sequencing. J Infect Dis. 2013 Feb;207(4):675–86.10.1093/infdis/jis734Search in Google Scholar PubMed PubMed Central
[36] Roxas BAP, Roxas JL, Claus-Walker R, Harishankar A, Mansoor A, Anwar F, et al. Phylogenomic analysis of Clostridioides difficile ribotype 106 strains reveals novel genetic islands and emergent phenotypes. Sci Rep. 2020 Dec;10(1):22135.10.1038/s41598-020-79123-2Search in Google Scholar PubMed PubMed Central
[37] O’Grady K, Riley TV, Knight DR. Complete genome assemblies of three highly prevalent, toxigenic Clostridioides difficile strains causing health care-associated infections in Australia. Rasko D, editor. Microbiol Resour Announc. 2021 Aug;10(31):e00599–21.10.1128/MRA.00599-21Search in Google Scholar PubMed PubMed Central
[38] Stabler RA, He M, Dawson L, Martin M, Valiente E, Corton C, et al. Comparative genome and phenotypic analysis of Clostridium difficile 027 strains provides insight into the evolution of a hypervirulent bacterium. Genome Biol. 2009;10(9):R102.10.1186/gb-2009-10-9-r102Search in Google Scholar PubMed PubMed Central
[39] Tettelin H, Riley D, Cattuto C, Medini D. Comparative genomics: the bacterial pan-genome. Curr Opin Microbiol. 2008 Oct;11(5):472–7.10.1016/j.mib.2008.09.006Search in Google Scholar PubMed
[40] Janoir C. Virulence factors of Clostridium difficile and their role during infection. Anaerobe. 2016 Feb;37:13–24.10.1016/j.anaerobe.2015.10.009Search in Google Scholar PubMed
[41] Squire MM, Carter GP, Mackin KE, Chakravorty A, Norén T, Elliott B, et al. Novel molecular type of Clostridium difficile in neonatal pigs, Western Australia. Emerg Infect Dis. 2013 May;19(5):790–2.10.3201/eid1905.121062Search in Google Scholar PubMed PubMed Central
[42] Goorhuis A, Bakker D, Corver J, Debast SB, Harmanus C, Notermans DW, et al. Emergence of Clostridium difficile infection due to a new hypervirulent strain, polymerase chain reaction ribotype 078. Clin Infect Dis Publ Infect Dis Soc Am. 2008 Nov;47(9):1162–70.10.1086/592257Search in Google Scholar PubMed
[43] Knight DR, Kullin B, Androga GO, Barbut F, Eckert C, Johnson S, et al. Evolutionary and genomic insights into Clostridioides difficile sequence type 11: a diverse zoonotic and antimicrobial-resistant lineage of global one health importance. Onderdonk AB, editor. mBio. 2019 Apr;10(2):e00446–19.10.1128/mBio.00446-19Search in Google Scholar PubMed PubMed Central
[44] Collins DA, Hawkey PM, Riley TV. Epidemiology of Clostridium difficile infection in Asia. Antimicrob Resist Infect Control. 2013;2(1):21.10.1186/2047-2994-2-21Search in Google Scholar PubMed PubMed Central
[45] Knight DR, Elliott B, Chang BJ, Perkins TT, Riley TV. Diversity and evolution in the genome of Clostridium difficile. Clin Microbiol Rev. 2015 Jul;28(3):721–41.10.1128/CMR.00127-14Search in Google Scholar PubMed PubMed Central
[46] Rodriguez C, Taminiau B, Van Broeck J, Delmée M, Daube G. Clostridium difficile in food and animals: a comprehensive review. In: Donelli G, editor. Advances in microbiology, infectious diseases and public health [Internet]. Cham: Springer International Publishing; 2016. p. 65–92. (Advances in Experimental Medicine and Biology; vol. 932). [cited 2024 Jan 6] https://link.springer.com/10.1007/5584_2016_27.10.1007/5584_2016_27Search in Google Scholar PubMed
[47] Jhung MA, Thompson AD, Killgore GE, Zukowski WE, Songer G, Warny M, et al. Toxinotype V Clostridium difficile in humans and food animals. Emerg Infect Dis. 2008 Jul;14(7):1039–45.10.3201/eid1407.071641Search in Google Scholar PubMed PubMed Central
[48] Bauer MP, Notermans DW, van Benthem BH, Brazier JS, Wilcox MH, Rupnik M, et al. Clostridium difficile infection in Europe: a hospital-based survey. Lancet. 2011 Jan;377(9759):63–73.10.1016/S0140-6736(10)61266-4Search in Google Scholar PubMed
[49] Eckert C, Emirian A, Le Monnier A, Cathala L, De Montclos H, Goret J, et al. Prevalence and pathogenicity of binary toxin-positive Clostridium difficile strains that do not produce toxins A and B. N Microbes N Infect. 2015 Jan;3:12–7.10.1016/j.nmni.2014.10.003Search in Google Scholar PubMed PubMed Central
[50] Zhang WZ, Wu Q, Li WG, Wang YY, Jia XX, Lu JX, et al. Analysis of molecular types and drug resistance characteristics of Clostridium difficile isolated from children under five years old with diarrhea. Dis Surveill. 2021;36(4):329–34.Search in Google Scholar
[51] Wang YY, Wang JR, Zheng WQ, Shen HM, Lv YY, Guo SF. Multilocus sequence typing of Shiga toxin genes in Clostridium difficile isolated from diarrhea patients in the Huhhot area. Clin Lab Med. 2021;36(11):1131–4.Search in Google Scholar
[52] Knight DR, Putsathit P, Elliott B, Riley TV. Contamination of Australian newborn calf carcasses at slaughter with Clostridium difficile. Clin Microbiol Infect Publ Eur Soc Clin Microbiol Infect Dis. 2016 Mar;22(3):266.e1–7.10.1016/j.cmi.2015.11.017Search in Google Scholar PubMed
[53] Elliott B, Dingle KE, Didelot X, Crook DW, Riley TV. The complexity and diversity of the pathogenicity locus in Clostridium difficile clade 5. Genome Biol Evol. 2014 Dec;6(12):3159–70.10.1093/gbe/evu248Search in Google Scholar PubMed PubMed Central
[54] Liu XS, Li WG, Zhang WZ, Wu Y, Lu JX. Molecular characterization of Clostridium difficile isolates in China from 2010 to 2015. Front Microbiol. 2018 Apr;9:845.10.3389/fmicb.2018.00845Search in Google Scholar PubMed PubMed Central
[55] Rupnik M, Wilcox MH, Gerding DN. Clostridium difficile infection: new developments in epidemiology and pathogenesis. Nat Rev Microbiol. 2009 Jul;7(7):526–36.10.1038/nrmicro2164Search in Google Scholar PubMed
© 2025 the author(s), published by De Gruyter
This work is licensed under the Creative Commons Attribution 4.0 International License.
Articles in the same Issue
- Safety assessment and modulation of hepatic CYP3A4 and UGT enzymes by Glycyrrhiza glabra aqueous extract in female Sprague–Dawley rats
- Adult-onset Still’s disease with hemophagocytic lymphohistiocytosis and minimal change disease
- Role of DZ2002 in reducing corneal graft rejection in rats by influencing Th17 activation via inhibition of the PI3K/AKT pathway and downregulation of TRAF1
- Biomedical Sciences
- Mechanism of triptolide regulating proliferation and apoptosis of hepatoma cells by inhibiting JAK/STAT pathway
- Maslinic acid improves mitochondrial function and inhibits oxidative stress and autophagy in human gastric smooth muscle cells
- Comparative analysis of inflammatory biomarkers for the diagnosis of neonatal sepsis: IL-6, IL-8, SAA, CRP, and PCT
- Post-pandemic insights on COVID-19 and premature ovarian insufficiency
- Proteome differences of dental stem cells between permanent and deciduous teeth by data-independent acquisition proteomics
- Optimizing a modified cetyltrimethylammonium bromide protocol for fungal DNA extraction: Insights from multilocus gene amplification
- Preliminary analysis of the role of small hepatitis B surface proteins mutations in the pathogenesis of occult hepatitis B infection via the endoplasmic reticulum stress-induced UPR-ERAD pathway
- Efficacy of alginate-coated gold nanoparticles against antibiotics-resistant Staphylococcus and Streptococcus pathogens of acne origins
- Battling COVID-19 leveraging nanobiotechnology: Gold and silver nanoparticle–B-escin conjugates as SARS-CoV-2 inhibitors
- Neurodegenerative diseases and neuroinflammation-induced apoptosis
- Impact of fracture fixation surgery on cognitive function and the gut microbiota in mice with a history of stroke
- COLEC10: A potential tumor suppressor and prognostic biomarker in hepatocellular carcinoma through modulation of EMT and PI3K-AKT pathways
- High-temperature requirement serine protease A2 inhibitor UCF-101 ameliorates damaged neurons in traumatic brain-injured rats by the AMPK/NF-κB pathway
- SIK1 inhibits IL-1β-stimulated cartilage apoptosis and inflammation in vitro through the CRTC2/CREB1 signaling
- Rutin–chitooligosaccharide complex: Comprehensive evaluation of its anti-inflammatory and analgesic properties in vitro and in vivo
- Knockdown of Aurora kinase B alleviates high glucose-triggered trophoblast cells damage and inflammation during gestational diabetes
- Calcium-sensing receptors promoted Homer1 expression and osteogenic differentiation in bone marrow mesenchymal stem cells
- ABI3BP can inhibit the proliferation, invasion, and epithelial–mesenchymal transition of non-small-cell lung cancer cells
- Changes in blood glucose and metabolism in hyperuricemia mice
- Rapid detection of the GJB2 c.235delC mutation based on CRISPR-Cas13a combined with lateral flow dipstick
- IL-11 promotes Ang II-induced autophagy inhibition and mitochondrial dysfunction in atrial fibroblasts
- Short-chain fatty acid attenuates intestinal inflammation by regulation of gut microbial composition in antibiotic-associated diarrhea
- Application of metagenomic next-generation sequencing in the diagnosis of pathogens in patients with diabetes complicated by community-acquired pneumonia
- NAT10 promotes radiotherapy resistance in non-small cell lung cancer by regulating KPNB1-mediated PD-L1 nuclear translocation
- Phytol-mixed micelles alleviate dexamethasone-induced osteoporosis in zebrafish: Activation of the MMP3–OPN–MAPK pathway-mediating bone remodeling
- Association between TGF-β1 and β-catenin expression in the vaginal wall of patients with pelvic organ prolapse
- Primary pleomorphic liposarcoma involving bilateral ovaries: Case report and literature review
- Effects of de novo donor-specific Class I and II antibodies on graft outcomes after liver transplantation: A pilot cohort study
- Sleep architecture in Alzheimer’s disease continuum: The deep sleep question
- Ephedra fragilis plant extract: A groundbreaking corrosion inhibitor for mild steel in acidic environments – electrochemical, EDX, DFT, and Monte Carlo studies
- Langerhans cell histiocytosis in an adult patient with upper jaw and pulmonary involvement: A case report
- Inhibition of mast cell activation by Jaranol-targeted Pirin ameliorates allergic responses in mouse allergic rhinitis
- Aeromonas veronii-induced septic arthritis of the hip in a child with acute lymphoblastic leukemia
- Clusterin activates the heat shock response via the PI3K/Akt pathway to protect cardiomyocytes from high-temperature-induced apoptosis
- Research progress on fecal microbiota transplantation in tumor prevention and treatment
- Low-pressure exposure influences the development of HAPE
- Stigmasterol alleviates endplate chondrocyte degeneration through inducing mitophagy by enhancing PINK1 mRNA acetylation via the ESR1/NAT10 axis
- AKAP12, mediated by transcription factor 21, inhibits cell proliferation, metastasis, and glycolysis in lung squamous cell carcinoma
- Association between PAX9 or MSX1 gene polymorphism and tooth agenesis risk: A meta-analysis
- A case of bloodstream infection caused by Neisseria gonorrhoeae
- Case of nasopharyngeal tuberculosis complicated with cervical lymph node and pulmonary tuberculosis
- p-Cymene inhibits pro-fibrotic and inflammatory mediators to prevent hepatic dysfunction
- GFPT2 promotes paclitaxel resistance in epithelial ovarian cancer cells via activating NF-κB signaling pathway
- Transfer RNA-derived fragment tRF-36 modulates varicose vein progression via human vascular smooth muscle cell Notch signaling
- RTA-408 attenuates the hepatic ischemia reperfusion injury in mice possibly by activating the Nrf2/HO-1 signaling pathway
- Decreased serum TIMP4 levels in patients with rheumatoid arthritis
- Sirt1 protects lupus nephritis by inhibiting the NLRP3 signaling pathway in human glomerular mesangial cells
- Sodium butyrate aids brain injury repair in neonatal rats
- Interaction of MTHFR polymorphism with PAX1 methylation in cervical cancer
- Convallatoxin inhibits proliferation and angiogenesis of glioma cells via regulating JAK/STAT3 pathway
- The effect of the PKR inhibitor, 2-aminopurine, on the replication of influenza A virus, and segment 8 mRNA splicing
- Effects of Ire1 gene on virulence and pathogenicity of Candida albicans
- Small cell lung cancer with small intestinal metastasis: Case report and literature review
- GRB14: A prognostic biomarker driving tumor progression in gastric cancer through the PI3K/AKT signaling pathway by interacting with COBLL1
- 15-Lipoxygenase-2 deficiency induces foam cell formation that can be restored by salidroside through the inhibition of arachidonic acid effects
- FTO alleviated the diabetic nephropathy progression by regulating the N6-methyladenosine levels of DACT1
- Clinical relevance of inflammatory markers in the evaluation of severity of ulcerative colitis: A retrospective study
- Zinc valproic acid complex promotes osteoblast differentiation and exhibits anti-osteoporotic potential
- Primary pulmonary synovial sarcoma in the bronchial cavity: A case report
- Metagenomic next-generation sequencing of alveolar lavage fluid improves the detection of pulmonary infection
- Uterine tumor resembling ovarian sex cord tumor with extensive rhabdoid differentiation: A case report
- Genomic analysis of a novel ST11(PR34365) Clostridioides difficile strain isolated from the human fecal of a CDI patient in Guizhou, China
- Effects of tiered cardiac rehabilitation on CRP, TNF-α, and physical endurance in older adults with coronary heart disease
- Changes in T-lymphocyte subpopulations in patients with colorectal cancer before and after acupoint catgut embedding acupuncture observation
- Modulating the tumor microenvironment: The role of traditional Chinese medicine in improving lung cancer treatment
- Alterations of metabolites related to microbiota–gut–brain axis in plasma of colon cancer, esophageal cancer, stomach cancer, and lung cancer patients
- Research on individualized drug sensitivity detection technology based on bio-3D printing technology for precision treatment of gastrointestinal stromal tumors
- CEBPB promotes ulcerative colitis-associated colorectal cancer by stimulating tumor growth and activating the NF-κB/STAT3 signaling pathway
- Oncolytic bacteria: A revolutionary approach to cancer therapy
- A de novo meningioma with rapid growth: A possible malignancy imposter?
- Diagnosis of secondary tuberculosis infection in an asymptomatic elderly with cancer using next-generation sequencing: Case report
- Hesperidin and its zinc(ii) complex enhance osteoblast differentiation and bone formation: In vitro and in vivo evaluations
- Research progress on the regulation of autophagy in cardiovascular diseases by chemokines
- Anti-arthritic, immunomodulatory, and inflammatory regulation by the benzimidazole derivative BMZ-AD: Insights from an FCA-induced rat model
- Immunoassay for pyruvate kinase M1/2 as an Alzheimer’s biomarker in CSF
- The role of HDAC11 in age-related hearing loss: Mechanisms and therapeutic implications
- Evaluation and application analysis of animal models of PIPNP based on data mining
- Therapeutic approaches for liver fibrosis/cirrhosis by targeting pyroptosis
- Fabrication of zinc oxide nanoparticles using Ruellia tuberosa leaf extract induces apoptosis through P53 and STAT3 signalling pathways in prostate cancer cells
- Haplo-hematopoietic stem cell transplantation and immunoradiotherapy for severe aplastic anemia complicated with nasopharyngeal carcinoma: A case report
- Modulation of the KEAP1-NRF2 pathway by Erianin: A novel approach to reduce psoriasiform inflammation and inflammatory signaling
- The expression of epidermal growth factor receptor 2 and its relationship with tumor-infiltrating lymphocytes and clinical pathological features in breast cancer patients
- Innovations in MALDI-TOF Mass Spectrometry: Bridging modern diagnostics and historical insights
- BAP1 complexes with YY1 and RBBP7 and its downstream targets in ccRCC cells
- Hypereosinophilic syndrome with elevated IgG4 and T-cell clonality: A report of two cases
- Electroacupuncture alleviates sciatic nerve injury in sciatica rats by regulating BDNF and NGF levels, myelin sheath degradation, and autophagy
- Polydatin prevents cholesterol gallstone formation by regulating cholesterol metabolism via PPAR-γ signaling
- RNF144A and RNF144B: Important molecules for health
- Analysis of the detection rate and related factors of thyroid nodules in the healthy population
- Artesunate inhibits hepatocellular carcinoma cell migration and invasion through OGA-mediated O-GlcNAcylation of ZEB1
- Endovascular management of post-pancreatectomy hemorrhage caused by a hepatic artery pseudoaneurysm: Case report and review of the literature
- Efficacy and safety of anti-PD-1/PD-L1 antibodies in patients with relapsed refractory diffuse large B-cell lymphoma: A meta-analysis
- SATB2 promotes humeral fracture healing in rats by activating the PI3K/AKT pathway
- Overexpression of the ferroptosis-related gene, NFS1, corresponds to gastric cancer growth and tumor immune infiltration
- Understanding risk factors and prognosis in diabetic foot ulcers
- Atractylenolide I alleviates the experimental allergic response in mice by suppressing TLR4/NF-kB/NLRP3 signalling
- FBXO31 inhibits the stemness characteristics of CD147 (+) melanoma stem cells
- Immune molecule diagnostics in colorectal cancer: CCL2 and CXCL11
- Inhibiting CXCR6 promotes senescence of activated hepatic stellate cells with limited proinflammatory SASP to attenuate hepatic fibrosis
- Cadmium toxicity, health risk and its remediation using low-cost biochar adsorbents
- Pulmonary cryptococcosis with headache as the first presentation: A case report
- Solitary pulmonary metastasis with cystic airspaces in colon cancer: A rare case report
- RUNX1 promotes denervation-induced muscle atrophy by activating the JUNB/NF-κB pathway and driving M1 macrophage polarization
- Morphometric analysis and immunobiological investigation of Indigofera oblongifolia on the infected lung with Plasmodium chabaudi
- The NuA4/TIP60 histone-modifying complex and Hr78 modulate the Lobe2 mutant eye phenotype
- Experimental study on salmon demineralized bone matrix loaded with recombinant human bone morphogenetic protein-2: In vitro and in vivo study
- A case of IgA nephropathy treated with a combination of telitacicept and half-dose glucocorticoids
- Analgesic and toxicological evaluation of cannabidiol-rich Moroccan Cannabis sativa L. (Khardala variety) extract: Evidence from an in vivo and in silico study
- Wound healing and signaling pathways
- Combination of immunotherapy and whole-brain radiotherapy on prognosis of patients with multiple brain metastases: A retrospective cohort study
- To explore the relationship between endometrial hyperemia and polycystic ovary syndrome
- Research progress on the impact of curcumin on immune responses in breast cancer
- Biogenic Cu/Ni nanotherapeutics from Descurainia sophia (L.) Webb ex Prantl seeds for the treatment of lung cancer
- Dapagliflozin attenuates atrial fibrosis via the HMGB1/RAGE pathway in atrial fibrillation rats
- Glycitein alleviates inflammation and apoptosis in keratinocytes via ROS-associated PI3K–Akt signalling pathway
- ADH5 inhibits proliferation but promotes EMT in non-small cell lung cancer cell through activating Smad2/Smad3
- Apoptotic efficacies of AgNPs formulated by Syzygium aromaticum leaf extract on 32D-FLT3-ITD human leukemia cell line with PI3K/AKT/mTOR signaling pathway
- Novel cuproptosis-related genes C1QBP and PFKP identified as prognostic and therapeutic targets in lung adenocarcinoma
- Bee venom promotes exosome secretion and alters miRNA cargo in T cells
- Treatment of pure red cell aplasia in a chronic kidney disease patient with roxadustat: A case report
- Comparative bioinformatics analysis of the Wnt pathway in breast cancer: Selection of novel biomarker panels associated with ER status
- Kynurenine facilitates renal cell carcinoma progression by suppressing M2 macrophage pyroptosis through inhibition of CASP1 cleavage
- RFX5 promotes the growth, motility, and inhibits apoptosis of gastric adenocarcinoma cells through the SIRT1/AMPK axis
- ALKBH5 exacerbates early cardiac damage after radiotherapy for breast cancer via m6A demethylation of TLR4
- Phytochemicals of Roman chamomile: Antioxidant, anti-aging, and whitening activities of distillation residues
- Circadian gene Cry1 inhibits the tumorigenicity of hepatocellular carcinoma by the BAX/BCL2-mediated apoptosis pathway
- The TNFR-RIPK1/RIPK3 signalling pathway mediates the effect of lanthanum on necroptosis of nerve cells
- Longitudinal monitoring of autoantibody dynamics in patients with early-stage non-small-cell lung cancer undergoing surgery
- The potential role of rutin, a flavonoid, in the management of cancer through modulation of cell signaling pathways
- Construction of pectinase gene engineering microbe and its application in tobacco sheets
- Construction of a microbial abundance prognostic scoring model based on intratumoral microbial data for predicting the prognosis of lung squamous cell carcinoma
- Sepsis complicated by haemophagocytic lymphohistiocytosis triggered by methicillin-resistant Staphylococcus aureus and human herpesvirus 8 in an immunocompromised elderly patient: A case report
- Sarcopenia in liver transplantation: A comprehensive bibliometric study of current research trends and future directions
- Advances in cancer immunotherapy and future directions in personalized medicine
- Can coronavirus disease 2019 affect male fertility or cause spontaneous abortion? A two-sample Mendelian randomization analysis
- Heat stroke associated with novel leukaemia inhibitory factor receptor gene variant in a Chinese infant
- PSME2 exacerbates ulcerative colitis by disrupting intestinal barrier function and promoting autophagy-dependent inflammation
- Hyperosmolar hyperglycemic state with severe hypernatremia coexisting with central diabetes insipidus: A case report and literature review
- Efficacy and mechanism of escin in improving the tissue microenvironment of blood vessel walls via anti-inflammatory and anticoagulant effects: Implications for clinical practice
- Merkel cell carcinoma: Clinicopathological analysis of three patients and literature review
- Genetic variants in VWF exon 26 and their implications for type 1 Von Willebrand disease in a Saudi Arabian population
- Lipoxin A4 improves myocardial ischemia/reperfusion injury through the Notch1-Nrf2 signaling pathway
- High levels of EPHB2 expression predict a poor prognosis and promote tumor progression in endometrial cancer
- Knockdown of SHP-2 delays renal tubular epithelial cell injury in diabetic nephropathy by inhibiting NLRP3 inflammasome-mediated pyroptosis
- Exploring the toxicity mechanisms and detoxification methods of Rhizoma Paridis
- Concomitant gastric carcinoma and primary hepatic angiosarcoma in a patient: A case report
- YAP1 inhibition protects retinal vascular endothelial cells under high glucose by inhibiting autophagy
- Identification of secretory protein related biomarkers for primary biliary cholangitis based on machine learning and experimental validation
- Integrated genomic and clinical modeling for prognostic assessment of radiotherapy response in rectal neoplasms
- Stem cell-based approaches for glaucoma treatment: a mini review
- Bacteriophage titering by optical density means: KOTE assays
- Neutrophil-related signature characterizes immune landscape and predicts prognosis of esophageal squamous cell carcinoma
- Integrated bioinformatic analysis and machine learning strategies to identify new potential immune biomarkers for Alzheimer’s disease and their targeting prediction with geniposide
- TRIM21 accelerates ferroptosis in intervertebral disc degeneration by promoting SLC7A11 ubiquitination and degradation
- TRIM21 accelerates ferroptosis in intervertebral disc degeneration by promoting SLC7A11 ubiquitination and degradation
- Histone modification and non-coding RNAs in skin aging: emerging therapeutic avenues
- A multiplicative behavioral model of DNA replication initiation in cells
- Biogenic gold nanoparticles synthesized from Pergularia daemia leaves: a novel approach for nasopharyngeal carcinoma therapy
- Creutzfeldt-Jakob disease mimicking Hashimoto’s encephalopathy: steroid response followed by decline
- Impact of semaphorin, Sema3F, on the gene transcription and protein expression of CREB and its binding protein CREBBP in primary hippocampal neurons of rats
- Iron overloaded M0 macrophages regulate hematopoietic stem cell proliferation and senescence via the Nrf2/Keap1/HO-1 pathway
- Revisiting the link between NADPH oxidase p22phox C242T polymorphism and ischemic stroke risk: an updated meta-analysis
- Exercise training preferentially modulates α1D-adrenergic receptor expression in peripheral arteries of hypertensive rats
- Overexpression of HE4/WFDC2 gene in mice leads to keratitis and corneal opacity
- Tumoral calcinosis complicating CKD-MBD in hemodialysis: a case report
- Mechanism of KLF4 Inhibition of epithelial-mesenchymal transition in gastric cancer cells
- Dissecting the molecular mechanisms of T cell infiltration in psoriatic lesions via cell-cell communication and regulatory network analysis
- Circadian rhythm-based prognostic features predict immune infiltration and tumor microenvironment in molecular subtypes of hepatocellular carcinoma
- Ecology and Environmental Science
- Optimization and comparative study of Bacillus consortia for cellulolytic potential and cellulase enzyme activity
- The complete mitochondrial genome analysis of Haemaphysalis hystricis Supino, 1897 (Ixodida: Ixodidae) and its phylogenetic implications
- Epidemiological characteristics and risk factors analysis of multidrug-resistant tuberculosis among tuberculosis population in Huzhou City, Eastern China
- Indices of human impacts on landscapes: How do they reflect the proportions of natural habitats?
- Genetic analysis of the Siberian flying squirrel population in the northern Changbai Mountains, Northeast China: Insights into population status and conservation
- Diversity and environmental drivers of Suillus communities in Pinus sylvestris var. mongolica forests of Inner Mongolia
- Global assessment of the fate of nitrogen deposition in forest ecosystems: Insights from 15N tracer studies
- Fungal and bacterial pathogenic co-infections mainly lead to the assembly of microbial community in tobacco stems
- Influencing of coal industry related airborne particulate matter on ocular surface tear film injury and inflammatory factor expression in Sprague-Dawley rats
- Temperature-dependent development, predation, and life table of Sphaerophoria macrogaster (Thomson) (Diptera: Syrphidae) feeding on Myzus persicae (Sulzer) (Homoptera: Aphididae)
- Eleonora’s falcon trophic interactions with insects within its breeding range: A systematic review
- Agriculture
- Integrated analysis of transcriptome, sRNAome, and degradome involved in the drought-response of maize Zhengdan958
- Variation in flower frost tolerance among seven apple cultivars and transcriptome response patterns in two contrastingly frost-tolerant selected cultivars
- Heritability of durable resistance to stripe rust in bread wheat (Triticum aestivum L.)
- Molecular mechanism of follicular development in laying hens based on the regulation of water metabolism
- Molecular identification and control studies on Coridius sp. (Hemiptera: Dinidoridae) in Al-Khamra, south of Jeddah, Saudi Arabia
- 10.1515/biol-2025-1218
- Animal Science
- Effect of sex ratio on the life history traits of an important invasive species, Spodoptera frugiperda
- Plant Sciences
- Hairpin in a haystack: In silico identification and characterization of plant-conserved microRNA in Rafflesiaceae
- Widely targeted metabolomics of different tissues in Rubus corchorifolius
- The complete chloroplast genome of Gerbera piloselloides (L.) Cass., 1820 (Carduoideae, Asteraceae) and its phylogenetic analysis
- Field trial to correlate mineral solubilization activity of Pseudomonas aeruginosa and biochemical content of groundnut plants
- Correlation analysis between semen routine parameters and sperm DNA fragmentation index in patients with semen non-liquefaction: A retrospective study
- Plasticity of the anatomical traits of Rhododendron L. (Ericaceae) leaves and its implications in adaptation to the plateau environment
- Effects of Piriformospora indica and arbuscular mycorrhizal fungus on growth and physiology of Moringa oleifera under low-temperature stress
- Effects of different sources of potassium fertiliser on yield, fruit quality and nutrient absorption in “Harward” kiwifruit (Actinidia deliciosa)
- Comparative efficiency and residue levels of spraying programs against powdery mildew in grape varieties
- The DREB7 transcription factor enhances salt tolerance in soybean plants under salt stress
- Using plant electrical signals of water hyacinth (Eichhornia crassipes) for water pollution monitoring
- Response of hybrid grapes (Vitis spp.) to two biotic stress factors and their seedlessness status
- Metabolomic profiling reveals systemic metabolic reprogramming in Alternaria alternata under salt stress
- Effects of mixed salinity and alkali stress on photosynthetic characteristics and PEPC gene expression of vegetable soybean seedlings
- Food Science
- Phytochemical analysis of Stachys iva: Discovering the optimal extract conditions and its bioactive compounds
- Review on role of honey in disease prevention and treatment through modulation of biological activities
- Computational analysis of polymorphic residues in maltose and maltotriose transporters of a wild Saccharomyces cerevisiae strain
- Optimization of phenolic compound extraction from Tunisian squash by-products: A sustainable approach for antioxidant and antibacterial applications
- Liupao tea aqueous extract alleviates dextran sulfate sodium-induced ulcerative colitis in rats by modulating the gut microbiota
- Toxicological qualities and detoxification trends of fruit by-products for valorization: A review
- Polyphenolic spectrum of cornelian cherry fruits and their health-promoting effect
- Optimizing the encapsulation of the refined extract of squash peels for functional food applications: A sustainable approach to reduce food waste
- Advancements in curcuminoid formulations: An update on bioavailability enhancement strategies curcuminoid bioavailability and formulations
- Impact of saline sprouting on antioxidant properties and bioactive compounds in chia seeds
- The dilemma of food genetics and improvement
- Causal effects of trace elements on congenital foot deformities and their subtypes: a Mendelian randomization study with gut microbiota mediation
- Honey meets acidity: a novel biopreservative approach against foodborne pathogens
- Bioengineering and Biotechnology
- Impact of hyaluronic acid-modified hafnium metalorganic frameworks containing rhynchophylline on Alzheimer’s disease
- Emerging patterns in nanoparticle-based therapeutic approaches for rheumatoid arthritis: A comprehensive bibliometric and visual analysis spanning two decades
- Application of CRISPR/Cas gene editing for infectious disease control in poultry
- Preparation of hafnium nitride-coated titanium implants by magnetron sputtering technology and evaluation of their antibacterial properties and biocompatibility
- Preparation and characterization of lemongrass oil nanoemulsion: Antimicrobial, antibiofilm, antioxidant, and anticancer activities
- Fluorescent detection of sialic acid–binding lectins using functionalized quantum dots in ELISA format
- Smart tectorigenin-loaded ZnO hydrogel nanocomposites for targeted wound healing: synthesis, characterization, and biological evaluation
- Corrigendum
- Corrigendum to “Utilization of convolutional neural networks to analyze microscopic images for high-throughput screening of mesenchymal stem cells”
- Corrigendum to “Effects of Ire1 gene on virulence and pathogenicity of Candida albicans”
- Retraction
- Retraction of “Down-regulation of miR-539 indicates poor prognosis in patients with pancreatic cancer”
Articles in the same Issue
- Safety assessment and modulation of hepatic CYP3A4 and UGT enzymes by Glycyrrhiza glabra aqueous extract in female Sprague–Dawley rats
- Adult-onset Still’s disease with hemophagocytic lymphohistiocytosis and minimal change disease
- Role of DZ2002 in reducing corneal graft rejection in rats by influencing Th17 activation via inhibition of the PI3K/AKT pathway and downregulation of TRAF1
- Biomedical Sciences
- Mechanism of triptolide regulating proliferation and apoptosis of hepatoma cells by inhibiting JAK/STAT pathway
- Maslinic acid improves mitochondrial function and inhibits oxidative stress and autophagy in human gastric smooth muscle cells
- Comparative analysis of inflammatory biomarkers for the diagnosis of neonatal sepsis: IL-6, IL-8, SAA, CRP, and PCT
- Post-pandemic insights on COVID-19 and premature ovarian insufficiency
- Proteome differences of dental stem cells between permanent and deciduous teeth by data-independent acquisition proteomics
- Optimizing a modified cetyltrimethylammonium bromide protocol for fungal DNA extraction: Insights from multilocus gene amplification
- Preliminary analysis of the role of small hepatitis B surface proteins mutations in the pathogenesis of occult hepatitis B infection via the endoplasmic reticulum stress-induced UPR-ERAD pathway
- Efficacy of alginate-coated gold nanoparticles against antibiotics-resistant Staphylococcus and Streptococcus pathogens of acne origins
- Battling COVID-19 leveraging nanobiotechnology: Gold and silver nanoparticle–B-escin conjugates as SARS-CoV-2 inhibitors
- Neurodegenerative diseases and neuroinflammation-induced apoptosis
- Impact of fracture fixation surgery on cognitive function and the gut microbiota in mice with a history of stroke
- COLEC10: A potential tumor suppressor and prognostic biomarker in hepatocellular carcinoma through modulation of EMT and PI3K-AKT pathways
- High-temperature requirement serine protease A2 inhibitor UCF-101 ameliorates damaged neurons in traumatic brain-injured rats by the AMPK/NF-κB pathway
- SIK1 inhibits IL-1β-stimulated cartilage apoptosis and inflammation in vitro through the CRTC2/CREB1 signaling
- Rutin–chitooligosaccharide complex: Comprehensive evaluation of its anti-inflammatory and analgesic properties in vitro and in vivo
- Knockdown of Aurora kinase B alleviates high glucose-triggered trophoblast cells damage and inflammation during gestational diabetes
- Calcium-sensing receptors promoted Homer1 expression and osteogenic differentiation in bone marrow mesenchymal stem cells
- ABI3BP can inhibit the proliferation, invasion, and epithelial–mesenchymal transition of non-small-cell lung cancer cells
- Changes in blood glucose and metabolism in hyperuricemia mice
- Rapid detection of the GJB2 c.235delC mutation based on CRISPR-Cas13a combined with lateral flow dipstick
- IL-11 promotes Ang II-induced autophagy inhibition and mitochondrial dysfunction in atrial fibroblasts
- Short-chain fatty acid attenuates intestinal inflammation by regulation of gut microbial composition in antibiotic-associated diarrhea
- Application of metagenomic next-generation sequencing in the diagnosis of pathogens in patients with diabetes complicated by community-acquired pneumonia
- NAT10 promotes radiotherapy resistance in non-small cell lung cancer by regulating KPNB1-mediated PD-L1 nuclear translocation
- Phytol-mixed micelles alleviate dexamethasone-induced osteoporosis in zebrafish: Activation of the MMP3–OPN–MAPK pathway-mediating bone remodeling
- Association between TGF-β1 and β-catenin expression in the vaginal wall of patients with pelvic organ prolapse
- Primary pleomorphic liposarcoma involving bilateral ovaries: Case report and literature review
- Effects of de novo donor-specific Class I and II antibodies on graft outcomes after liver transplantation: A pilot cohort study
- Sleep architecture in Alzheimer’s disease continuum: The deep sleep question
- Ephedra fragilis plant extract: A groundbreaking corrosion inhibitor for mild steel in acidic environments – electrochemical, EDX, DFT, and Monte Carlo studies
- Langerhans cell histiocytosis in an adult patient with upper jaw and pulmonary involvement: A case report
- Inhibition of mast cell activation by Jaranol-targeted Pirin ameliorates allergic responses in mouse allergic rhinitis
- Aeromonas veronii-induced septic arthritis of the hip in a child with acute lymphoblastic leukemia
- Clusterin activates the heat shock response via the PI3K/Akt pathway to protect cardiomyocytes from high-temperature-induced apoptosis
- Research progress on fecal microbiota transplantation in tumor prevention and treatment
- Low-pressure exposure influences the development of HAPE
- Stigmasterol alleviates endplate chondrocyte degeneration through inducing mitophagy by enhancing PINK1 mRNA acetylation via the ESR1/NAT10 axis
- AKAP12, mediated by transcription factor 21, inhibits cell proliferation, metastasis, and glycolysis in lung squamous cell carcinoma
- Association between PAX9 or MSX1 gene polymorphism and tooth agenesis risk: A meta-analysis
- A case of bloodstream infection caused by Neisseria gonorrhoeae
- Case of nasopharyngeal tuberculosis complicated with cervical lymph node and pulmonary tuberculosis
- p-Cymene inhibits pro-fibrotic and inflammatory mediators to prevent hepatic dysfunction
- GFPT2 promotes paclitaxel resistance in epithelial ovarian cancer cells via activating NF-κB signaling pathway
- Transfer RNA-derived fragment tRF-36 modulates varicose vein progression via human vascular smooth muscle cell Notch signaling
- RTA-408 attenuates the hepatic ischemia reperfusion injury in mice possibly by activating the Nrf2/HO-1 signaling pathway
- Decreased serum TIMP4 levels in patients with rheumatoid arthritis
- Sirt1 protects lupus nephritis by inhibiting the NLRP3 signaling pathway in human glomerular mesangial cells
- Sodium butyrate aids brain injury repair in neonatal rats
- Interaction of MTHFR polymorphism with PAX1 methylation in cervical cancer
- Convallatoxin inhibits proliferation and angiogenesis of glioma cells via regulating JAK/STAT3 pathway
- The effect of the PKR inhibitor, 2-aminopurine, on the replication of influenza A virus, and segment 8 mRNA splicing
- Effects of Ire1 gene on virulence and pathogenicity of Candida albicans
- Small cell lung cancer with small intestinal metastasis: Case report and literature review
- GRB14: A prognostic biomarker driving tumor progression in gastric cancer through the PI3K/AKT signaling pathway by interacting with COBLL1
- 15-Lipoxygenase-2 deficiency induces foam cell formation that can be restored by salidroside through the inhibition of arachidonic acid effects
- FTO alleviated the diabetic nephropathy progression by regulating the N6-methyladenosine levels of DACT1
- Clinical relevance of inflammatory markers in the evaluation of severity of ulcerative colitis: A retrospective study
- Zinc valproic acid complex promotes osteoblast differentiation and exhibits anti-osteoporotic potential
- Primary pulmonary synovial sarcoma in the bronchial cavity: A case report
- Metagenomic next-generation sequencing of alveolar lavage fluid improves the detection of pulmonary infection
- Uterine tumor resembling ovarian sex cord tumor with extensive rhabdoid differentiation: A case report
- Genomic analysis of a novel ST11(PR34365) Clostridioides difficile strain isolated from the human fecal of a CDI patient in Guizhou, China
- Effects of tiered cardiac rehabilitation on CRP, TNF-α, and physical endurance in older adults with coronary heart disease
- Changes in T-lymphocyte subpopulations in patients with colorectal cancer before and after acupoint catgut embedding acupuncture observation
- Modulating the tumor microenvironment: The role of traditional Chinese medicine in improving lung cancer treatment
- Alterations of metabolites related to microbiota–gut–brain axis in plasma of colon cancer, esophageal cancer, stomach cancer, and lung cancer patients
- Research on individualized drug sensitivity detection technology based on bio-3D printing technology for precision treatment of gastrointestinal stromal tumors
- CEBPB promotes ulcerative colitis-associated colorectal cancer by stimulating tumor growth and activating the NF-κB/STAT3 signaling pathway
- Oncolytic bacteria: A revolutionary approach to cancer therapy
- A de novo meningioma with rapid growth: A possible malignancy imposter?
- Diagnosis of secondary tuberculosis infection in an asymptomatic elderly with cancer using next-generation sequencing: Case report
- Hesperidin and its zinc(ii) complex enhance osteoblast differentiation and bone formation: In vitro and in vivo evaluations
- Research progress on the regulation of autophagy in cardiovascular diseases by chemokines
- Anti-arthritic, immunomodulatory, and inflammatory regulation by the benzimidazole derivative BMZ-AD: Insights from an FCA-induced rat model
- Immunoassay for pyruvate kinase M1/2 as an Alzheimer’s biomarker in CSF
- The role of HDAC11 in age-related hearing loss: Mechanisms and therapeutic implications
- Evaluation and application analysis of animal models of PIPNP based on data mining
- Therapeutic approaches for liver fibrosis/cirrhosis by targeting pyroptosis
- Fabrication of zinc oxide nanoparticles using Ruellia tuberosa leaf extract induces apoptosis through P53 and STAT3 signalling pathways in prostate cancer cells
- Haplo-hematopoietic stem cell transplantation and immunoradiotherapy for severe aplastic anemia complicated with nasopharyngeal carcinoma: A case report
- Modulation of the KEAP1-NRF2 pathway by Erianin: A novel approach to reduce psoriasiform inflammation and inflammatory signaling
- The expression of epidermal growth factor receptor 2 and its relationship with tumor-infiltrating lymphocytes and clinical pathological features in breast cancer patients
- Innovations in MALDI-TOF Mass Spectrometry: Bridging modern diagnostics and historical insights
- BAP1 complexes with YY1 and RBBP7 and its downstream targets in ccRCC cells
- Hypereosinophilic syndrome with elevated IgG4 and T-cell clonality: A report of two cases
- Electroacupuncture alleviates sciatic nerve injury in sciatica rats by regulating BDNF and NGF levels, myelin sheath degradation, and autophagy
- Polydatin prevents cholesterol gallstone formation by regulating cholesterol metabolism via PPAR-γ signaling
- RNF144A and RNF144B: Important molecules for health
- Analysis of the detection rate and related factors of thyroid nodules in the healthy population
- Artesunate inhibits hepatocellular carcinoma cell migration and invasion through OGA-mediated O-GlcNAcylation of ZEB1
- Endovascular management of post-pancreatectomy hemorrhage caused by a hepatic artery pseudoaneurysm: Case report and review of the literature
- Efficacy and safety of anti-PD-1/PD-L1 antibodies in patients with relapsed refractory diffuse large B-cell lymphoma: A meta-analysis
- SATB2 promotes humeral fracture healing in rats by activating the PI3K/AKT pathway
- Overexpression of the ferroptosis-related gene, NFS1, corresponds to gastric cancer growth and tumor immune infiltration
- Understanding risk factors and prognosis in diabetic foot ulcers
- Atractylenolide I alleviates the experimental allergic response in mice by suppressing TLR4/NF-kB/NLRP3 signalling
- FBXO31 inhibits the stemness characteristics of CD147 (+) melanoma stem cells
- Immune molecule diagnostics in colorectal cancer: CCL2 and CXCL11
- Inhibiting CXCR6 promotes senescence of activated hepatic stellate cells with limited proinflammatory SASP to attenuate hepatic fibrosis
- Cadmium toxicity, health risk and its remediation using low-cost biochar adsorbents
- Pulmonary cryptococcosis with headache as the first presentation: A case report
- Solitary pulmonary metastasis with cystic airspaces in colon cancer: A rare case report
- RUNX1 promotes denervation-induced muscle atrophy by activating the JUNB/NF-κB pathway and driving M1 macrophage polarization
- Morphometric analysis and immunobiological investigation of Indigofera oblongifolia on the infected lung with Plasmodium chabaudi
- The NuA4/TIP60 histone-modifying complex and Hr78 modulate the Lobe2 mutant eye phenotype
- Experimental study on salmon demineralized bone matrix loaded with recombinant human bone morphogenetic protein-2: In vitro and in vivo study
- A case of IgA nephropathy treated with a combination of telitacicept and half-dose glucocorticoids
- Analgesic and toxicological evaluation of cannabidiol-rich Moroccan Cannabis sativa L. (Khardala variety) extract: Evidence from an in vivo and in silico study
- Wound healing and signaling pathways
- Combination of immunotherapy and whole-brain radiotherapy on prognosis of patients with multiple brain metastases: A retrospective cohort study
- To explore the relationship between endometrial hyperemia and polycystic ovary syndrome
- Research progress on the impact of curcumin on immune responses in breast cancer
- Biogenic Cu/Ni nanotherapeutics from Descurainia sophia (L.) Webb ex Prantl seeds for the treatment of lung cancer
- Dapagliflozin attenuates atrial fibrosis via the HMGB1/RAGE pathway in atrial fibrillation rats
- Glycitein alleviates inflammation and apoptosis in keratinocytes via ROS-associated PI3K–Akt signalling pathway
- ADH5 inhibits proliferation but promotes EMT in non-small cell lung cancer cell through activating Smad2/Smad3
- Apoptotic efficacies of AgNPs formulated by Syzygium aromaticum leaf extract on 32D-FLT3-ITD human leukemia cell line with PI3K/AKT/mTOR signaling pathway
- Novel cuproptosis-related genes C1QBP and PFKP identified as prognostic and therapeutic targets in lung adenocarcinoma
- Bee venom promotes exosome secretion and alters miRNA cargo in T cells
- Treatment of pure red cell aplasia in a chronic kidney disease patient with roxadustat: A case report
- Comparative bioinformatics analysis of the Wnt pathway in breast cancer: Selection of novel biomarker panels associated with ER status
- Kynurenine facilitates renal cell carcinoma progression by suppressing M2 macrophage pyroptosis through inhibition of CASP1 cleavage
- RFX5 promotes the growth, motility, and inhibits apoptosis of gastric adenocarcinoma cells through the SIRT1/AMPK axis
- ALKBH5 exacerbates early cardiac damage after radiotherapy for breast cancer via m6A demethylation of TLR4
- Phytochemicals of Roman chamomile: Antioxidant, anti-aging, and whitening activities of distillation residues
- Circadian gene Cry1 inhibits the tumorigenicity of hepatocellular carcinoma by the BAX/BCL2-mediated apoptosis pathway
- The TNFR-RIPK1/RIPK3 signalling pathway mediates the effect of lanthanum on necroptosis of nerve cells
- Longitudinal monitoring of autoantibody dynamics in patients with early-stage non-small-cell lung cancer undergoing surgery
- The potential role of rutin, a flavonoid, in the management of cancer through modulation of cell signaling pathways
- Construction of pectinase gene engineering microbe and its application in tobacco sheets
- Construction of a microbial abundance prognostic scoring model based on intratumoral microbial data for predicting the prognosis of lung squamous cell carcinoma
- Sepsis complicated by haemophagocytic lymphohistiocytosis triggered by methicillin-resistant Staphylococcus aureus and human herpesvirus 8 in an immunocompromised elderly patient: A case report
- Sarcopenia in liver transplantation: A comprehensive bibliometric study of current research trends and future directions
- Advances in cancer immunotherapy and future directions in personalized medicine
- Can coronavirus disease 2019 affect male fertility or cause spontaneous abortion? A two-sample Mendelian randomization analysis
- Heat stroke associated with novel leukaemia inhibitory factor receptor gene variant in a Chinese infant
- PSME2 exacerbates ulcerative colitis by disrupting intestinal barrier function and promoting autophagy-dependent inflammation
- Hyperosmolar hyperglycemic state with severe hypernatremia coexisting with central diabetes insipidus: A case report and literature review
- Efficacy and mechanism of escin in improving the tissue microenvironment of blood vessel walls via anti-inflammatory and anticoagulant effects: Implications for clinical practice
- Merkel cell carcinoma: Clinicopathological analysis of three patients and literature review
- Genetic variants in VWF exon 26 and their implications for type 1 Von Willebrand disease in a Saudi Arabian population
- Lipoxin A4 improves myocardial ischemia/reperfusion injury through the Notch1-Nrf2 signaling pathway
- High levels of EPHB2 expression predict a poor prognosis and promote tumor progression in endometrial cancer
- Knockdown of SHP-2 delays renal tubular epithelial cell injury in diabetic nephropathy by inhibiting NLRP3 inflammasome-mediated pyroptosis
- Exploring the toxicity mechanisms and detoxification methods of Rhizoma Paridis
- Concomitant gastric carcinoma and primary hepatic angiosarcoma in a patient: A case report
- YAP1 inhibition protects retinal vascular endothelial cells under high glucose by inhibiting autophagy
- Identification of secretory protein related biomarkers for primary biliary cholangitis based on machine learning and experimental validation
- Integrated genomic and clinical modeling for prognostic assessment of radiotherapy response in rectal neoplasms
- Stem cell-based approaches for glaucoma treatment: a mini review
- Bacteriophage titering by optical density means: KOTE assays
- Neutrophil-related signature characterizes immune landscape and predicts prognosis of esophageal squamous cell carcinoma
- Integrated bioinformatic analysis and machine learning strategies to identify new potential immune biomarkers for Alzheimer’s disease and their targeting prediction with geniposide
- TRIM21 accelerates ferroptosis in intervertebral disc degeneration by promoting SLC7A11 ubiquitination and degradation
- TRIM21 accelerates ferroptosis in intervertebral disc degeneration by promoting SLC7A11 ubiquitination and degradation
- Histone modification and non-coding RNAs in skin aging: emerging therapeutic avenues
- A multiplicative behavioral model of DNA replication initiation in cells
- Biogenic gold nanoparticles synthesized from Pergularia daemia leaves: a novel approach for nasopharyngeal carcinoma therapy
- Creutzfeldt-Jakob disease mimicking Hashimoto’s encephalopathy: steroid response followed by decline
- Impact of semaphorin, Sema3F, on the gene transcription and protein expression of CREB and its binding protein CREBBP in primary hippocampal neurons of rats
- Iron overloaded M0 macrophages regulate hematopoietic stem cell proliferation and senescence via the Nrf2/Keap1/HO-1 pathway
- Revisiting the link between NADPH oxidase p22phox C242T polymorphism and ischemic stroke risk: an updated meta-analysis
- Exercise training preferentially modulates α1D-adrenergic receptor expression in peripheral arteries of hypertensive rats
- Overexpression of HE4/WFDC2 gene in mice leads to keratitis and corneal opacity
- Tumoral calcinosis complicating CKD-MBD in hemodialysis: a case report
- Mechanism of KLF4 Inhibition of epithelial-mesenchymal transition in gastric cancer cells
- Dissecting the molecular mechanisms of T cell infiltration in psoriatic lesions via cell-cell communication and regulatory network analysis
- Circadian rhythm-based prognostic features predict immune infiltration and tumor microenvironment in molecular subtypes of hepatocellular carcinoma
- Ecology and Environmental Science
- Optimization and comparative study of Bacillus consortia for cellulolytic potential and cellulase enzyme activity
- The complete mitochondrial genome analysis of Haemaphysalis hystricis Supino, 1897 (Ixodida: Ixodidae) and its phylogenetic implications
- Epidemiological characteristics and risk factors analysis of multidrug-resistant tuberculosis among tuberculosis population in Huzhou City, Eastern China
- Indices of human impacts on landscapes: How do they reflect the proportions of natural habitats?
- Genetic analysis of the Siberian flying squirrel population in the northern Changbai Mountains, Northeast China: Insights into population status and conservation
- Diversity and environmental drivers of Suillus communities in Pinus sylvestris var. mongolica forests of Inner Mongolia
- Global assessment of the fate of nitrogen deposition in forest ecosystems: Insights from 15N tracer studies
- Fungal and bacterial pathogenic co-infections mainly lead to the assembly of microbial community in tobacco stems
- Influencing of coal industry related airborne particulate matter on ocular surface tear film injury and inflammatory factor expression in Sprague-Dawley rats
- Temperature-dependent development, predation, and life table of Sphaerophoria macrogaster (Thomson) (Diptera: Syrphidae) feeding on Myzus persicae (Sulzer) (Homoptera: Aphididae)
- Eleonora’s falcon trophic interactions with insects within its breeding range: A systematic review
- Agriculture
- Integrated analysis of transcriptome, sRNAome, and degradome involved in the drought-response of maize Zhengdan958
- Variation in flower frost tolerance among seven apple cultivars and transcriptome response patterns in two contrastingly frost-tolerant selected cultivars
- Heritability of durable resistance to stripe rust in bread wheat (Triticum aestivum L.)
- Molecular mechanism of follicular development in laying hens based on the regulation of water metabolism
- Molecular identification and control studies on Coridius sp. (Hemiptera: Dinidoridae) in Al-Khamra, south of Jeddah, Saudi Arabia
- 10.1515/biol-2025-1218
- Animal Science
- Effect of sex ratio on the life history traits of an important invasive species, Spodoptera frugiperda
- Plant Sciences
- Hairpin in a haystack: In silico identification and characterization of plant-conserved microRNA in Rafflesiaceae
- Widely targeted metabolomics of different tissues in Rubus corchorifolius
- The complete chloroplast genome of Gerbera piloselloides (L.) Cass., 1820 (Carduoideae, Asteraceae) and its phylogenetic analysis
- Field trial to correlate mineral solubilization activity of Pseudomonas aeruginosa and biochemical content of groundnut plants
- Correlation analysis between semen routine parameters and sperm DNA fragmentation index in patients with semen non-liquefaction: A retrospective study
- Plasticity of the anatomical traits of Rhododendron L. (Ericaceae) leaves and its implications in adaptation to the plateau environment
- Effects of Piriformospora indica and arbuscular mycorrhizal fungus on growth and physiology of Moringa oleifera under low-temperature stress
- Effects of different sources of potassium fertiliser on yield, fruit quality and nutrient absorption in “Harward” kiwifruit (Actinidia deliciosa)
- Comparative efficiency and residue levels of spraying programs against powdery mildew in grape varieties
- The DREB7 transcription factor enhances salt tolerance in soybean plants under salt stress
- Using plant electrical signals of water hyacinth (Eichhornia crassipes) for water pollution monitoring
- Response of hybrid grapes (Vitis spp.) to two biotic stress factors and their seedlessness status
- Metabolomic profiling reveals systemic metabolic reprogramming in Alternaria alternata under salt stress
- Effects of mixed salinity and alkali stress on photosynthetic characteristics and PEPC gene expression of vegetable soybean seedlings
- Food Science
- Phytochemical analysis of Stachys iva: Discovering the optimal extract conditions and its bioactive compounds
- Review on role of honey in disease prevention and treatment through modulation of biological activities
- Computational analysis of polymorphic residues in maltose and maltotriose transporters of a wild Saccharomyces cerevisiae strain
- Optimization of phenolic compound extraction from Tunisian squash by-products: A sustainable approach for antioxidant and antibacterial applications
- Liupao tea aqueous extract alleviates dextran sulfate sodium-induced ulcerative colitis in rats by modulating the gut microbiota
- Toxicological qualities and detoxification trends of fruit by-products for valorization: A review
- Polyphenolic spectrum of cornelian cherry fruits and their health-promoting effect
- Optimizing the encapsulation of the refined extract of squash peels for functional food applications: A sustainable approach to reduce food waste
- Advancements in curcuminoid formulations: An update on bioavailability enhancement strategies curcuminoid bioavailability and formulations
- Impact of saline sprouting on antioxidant properties and bioactive compounds in chia seeds
- The dilemma of food genetics and improvement
- Causal effects of trace elements on congenital foot deformities and their subtypes: a Mendelian randomization study with gut microbiota mediation
- Honey meets acidity: a novel biopreservative approach against foodborne pathogens
- Bioengineering and Biotechnology
- Impact of hyaluronic acid-modified hafnium metalorganic frameworks containing rhynchophylline on Alzheimer’s disease
- Emerging patterns in nanoparticle-based therapeutic approaches for rheumatoid arthritis: A comprehensive bibliometric and visual analysis spanning two decades
- Application of CRISPR/Cas gene editing for infectious disease control in poultry
- Preparation of hafnium nitride-coated titanium implants by magnetron sputtering technology and evaluation of their antibacterial properties and biocompatibility
- Preparation and characterization of lemongrass oil nanoemulsion: Antimicrobial, antibiofilm, antioxidant, and anticancer activities
- Fluorescent detection of sialic acid–binding lectins using functionalized quantum dots in ELISA format
- Smart tectorigenin-loaded ZnO hydrogel nanocomposites for targeted wound healing: synthesis, characterization, and biological evaluation
- Corrigendum
- Corrigendum to “Utilization of convolutional neural networks to analyze microscopic images for high-throughput screening of mesenchymal stem cells”
- Corrigendum to “Effects of Ire1 gene on virulence and pathogenicity of Candida albicans”
- Retraction
- Retraction of “Down-regulation of miR-539 indicates poor prognosis in patients with pancreatic cancer”

