Abstract
C24H31NO5, orthorhombic, P212121 (no. 19), a = 4.9020(10) Å, b = 17.638(4) Å, c = 24.848(5) Å, V = 2148.4(7) Å3, Z = 4, R gt (F) = 0.0751, wRref (F 2) = 0.1901, T = 293(2) K.
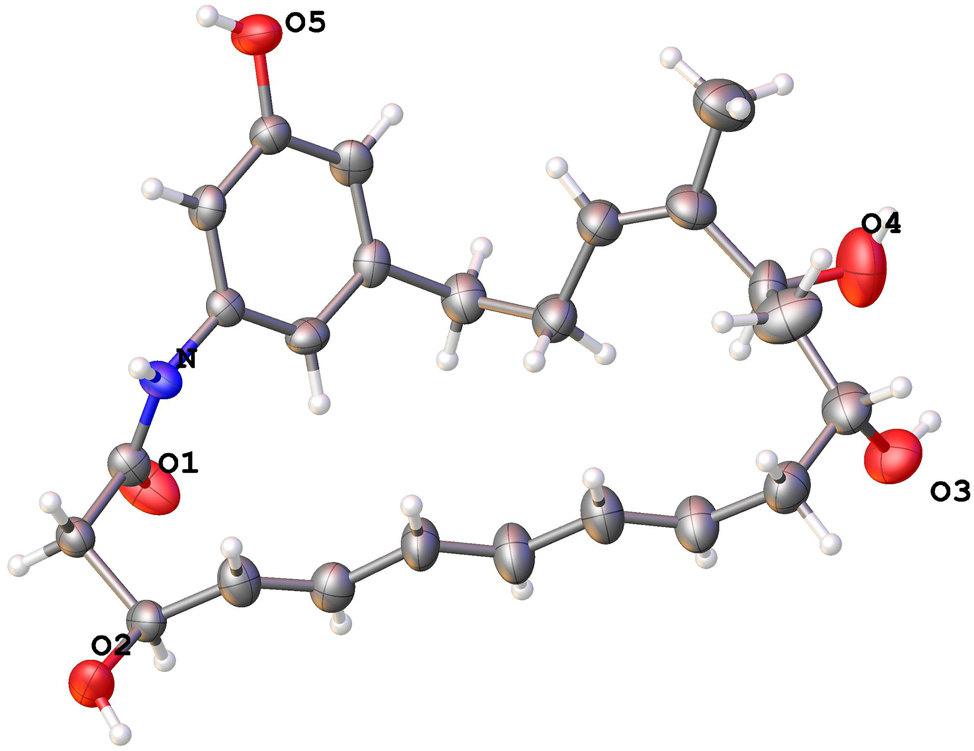
The molecular structure is shown in the figure. Table 1 contains the crystallographic data.
Data collection and handling.
| Crystal: | Colourless block |
| Size: | 0.20 × 0.10 × 0.05 mm |
| Wavelength: | MoKα radiation (0.71073 Å) |
| μ: | 0.09 mm−1 |
| Diffractometer, scan mode: | Nonius CAD-4, φ and ω scans |
| θ max, completeness: | 25.3°, 100 % |
| N(hkl)measured, N(hkl)unique, R int: | 4485, 3881, 0.078 |
| Criterion for I obs, N(hkl)gt: | I obs > 2 σ(I obs), 2112 |
| N(param)refined: | 264 |
| Programs: | Nonius, Olex2, 1 , 2 SHELX 3 , 4 |
1 Source of material
The title compound is a natural product, obtained by extraction and isolation from actinomycetes fermentation products. 5
2 Experimental details
The structure was treated with the Olex2 crystallographic software package, 1 , 2 solved with the SHELXT structure solution program and refined with the SHELXL 3 , 4 refinement package. Carbon-bound hydrogen atoms were placed in calculated positions and refined with riding coordinates, with U iso(H) fixed at 1.2 times of U eq(C) (Table 1).
3 Comment
Ansamycins are a family of broad-spectrum bioactive macrolactams, such as antituberculosis rifamycin, 5 antitumor maytansine 6 , 7 and Hsp90 inhibitor geldanamycin. 8 The title compound of C24H31NO5, named demethyltrienomycinol, has been reported in 2008, but no three-dimensional coordinates were deposited with the Cambridge Crystallographic Data Centre in the reported literature. 5 , 6 , 7 The structure of C24H31NO5 is composed of a 21-membered macrocyclic lactam ring including a triene moiety, which formed through an amide linkage to the 3-amino-5-hydroxybenzoic acid (AHBA) fragment. The title compound crystallizes in the orthorhombic space group P212121. The geometry of title structure was characterized with the bond angles and lengths in accordance with the literature. 8 In particular, the torsion angles of C2⋯C3⋯C4⋯C5, C21⋯N⋯C1⋯C2, O2⋯C3⋯C4⋯C5, C9⋯C10⋯C11⋯O3, O4⋯C13⋯C14⋯C15 and C16⋯C17⋯C18⋯C19, are −113.9(6)°, 169.9(4)°, 126 (3)°, −69.5(6)°, −56.2(7)° and −70.9(6)°, respectively. Furthermore, the bond lengths of O1⋯C1, O2⋯C35, C4⋯C5, O5⋯C23, C10⋯C11,C14⋯C16 and N⋯C1, are 1.248(6) Å, 1.428(6) Å, 1.330(7) Å, 1.359(6) Å, 1.501(8) Å, 1.313(7) Å, and 1.332(6) Å, coincident with the reported parameters. 9 And it was further unequivocally secured that the stereo configurations of C3, C11 and C13 are S, R and S, respectively. 10 , 11 In the crystal, the molecules are arranged in layers via hydrogen bonds O5–H⋯O2 with max D–A distance 2.9 Å and minimum angle 120°. All bond angles and lengths of the title structure are within the expected range.
Acknowledgements
This work was supported by the education natural science key project of Anhui Provice (2024AH051755), and the open research project of the rural revitalization collaborative technical service center of Anhui province (Huangshan University) (XCZXZD2402).
-
Author contributions: All the authors have accepted responsibility for the entire content of this submitted manuscript and approved submission.
-
Conflict of interest: The authors declare no conflicts of interest regarding this article.
-
Research funding: This work was supported by the education natural science key project of Anhui Provice (2024AH051755), and the open research project of the rural revitalization collaborative technical service center of Anhui province (Huangshan University) (XCZXZD2402).
References
1. Dolomanov, O. V.; Bourhis, L. J.; Gildea, R. J.; Howard, J. A. K.; Puschmann, H. Olex2: A Complete Structure Solution, Refinement and Analysis Program. J. Appl. Crystallogr. 2009, 42, 339–341; https://doi.org/10.1107/s0021889808042726.Suche in Google Scholar
2. Bourhis, L. J.; Dolomanov, O. V.; Gildea, R. J.; Howard, J. A. K.; Puschmann, H. The Anatomy of a Comprehensive Constrained, Restrained Refinement Program for the Modern computingenvironment–Olex2 Dissected. Acta Crystallogr. 2015, 71, 59–75; https://doi.org/10.1107/s2053273314022207.Suche in Google Scholar
3. Sheldrick, G. M. A Short History of Shelx. Acta Crystallogr. 2008, 64, 112–122; https://doi.org/10.1107/s0108767307043930.Suche in Google Scholar PubMed
4. Sheldrick, G. M. Crystal Structure Refinement with Shelxl. Acta Crystallogr. 2015, 71, 3–8; https://doi.org/10.1107/s2053229614024218.Suche in Google Scholar
5. Kawamura, T.; Tashiro, E.; Yamamoto, K.; Shindo, K.; Imoto, M. SAR Study of a Novel Triene-ansamycin Group Compound, Quinotrierixin and Related Compounds, as Inhibitors of Erstress-Induced xbp1 Activation. J. Antibiot. 2008, 61, 303–311; https://doi.org/10.1038/ja.2008.44.Suche in Google Scholar PubMed
6. Maggi, N.; Pasqualucci, C.; Ballotta, R.; Sensi, P. Rifampicin: A New Orally Active Rifamycin. Chemotherapy 1966, 11, 285–292; https://doi.org/10.1159/000220462.Suche in Google Scholar PubMed
7. Cassady, J. M.; Chan, K. K.; Floss, H. G.; Leistner, E. Recent Developments in the Maytansinoid Antitumor Agents. Chem. Pharm. Bull. 2004, 52, 1–26; https://doi.org/10.1248/cpb.52.1.Suche in Google Scholar PubMed
8. Kusari, P.; Kusari, S.; Eckelmann, D.; Zuhlke, S.; Kayser, O.; Spiteller, M. Cross-Species Biosynthesis of Maytansine in Maytenus Serrata. RSC Adv. 2016, 6, 10011–10016; https://doi.org/10.1039/c5ra25042k.Suche in Google Scholar
9. Yamada, Y.; Tashiro, E.; Taketani, S.; Imoto, M.; Kataoka, T. Mycotrienin Ii, A Translation Inhibitor that Prevents Icam-1 Expression Induced by Pro-Inflammatory Cytokines. J. Antibiot. 2011, 64, 361–366; https://doi.org/10.1038/ja.2011.23.Suche in Google Scholar PubMed
10. Whitesell, L.; Mimnaugh, E. G.; Costa, B. D.; Neckers, M. L. M.; Neckers, L. M. Inhibition of Heat Shock Protein HSP90-pp60v-src Heteroprotein Complex Formation by Benzoquinone Ansamycins: Essential Role for Stress Proteins in Oncogenic Transformation. Proc. Natl. Acad. Sci. U. S. A. 1994, 91, 8324–8328; https://doi.org/10.1073/pnas.91.18.8324.Suche in Google Scholar PubMed PubMed Central
11. Sun, W.; Li, Y. P.; Jin, J.; Li, Z. R.; Shan, G. Z. Crystal Structure of 17-(2′-(1′-oxa-4′-aza-heterocyclohexyl-1′-)Ethylamino)-17-Demethoxygeldanamycin Dihydrate, C34H54N4O11. Z. Kristallogr.-N. Cryst. Struct. 2015, 230, 180–182; https://doi.org/10.1515/ncrs-2014-9073.Suche in Google Scholar
© 2025 the author(s), published by De Gruyter, Berlin/Boston
This work is licensed under the Creative Commons Attribution 4.0 International License.
Artikel in diesem Heft
- Frontmatter
- New Crystal Structures
- The crystal structure of ( E )-4-chloro-7-(diethylamino)-3-(3-oxo-3-phenylprop-1-en-1-yl)-2 H -chromen-2-one C22H20ClNO3
- Crystal structure of (5S,6E,8E,10E,13R,15S,16Z)-15,5,13,15-tetrahydroxy-16-methyl-2-aza-1(1,3)-benzenacyclononadecaphane-6,8,10,16-tetraen-3-one, C24H31NO5
- The crystal structure of [bis(6-phenylpyridine-2-carboxylato-κ2O,N)-dihydrate-nickel(II)] ethanol solvate
- The crystal structure of poly[(μ3-2-nitro-1,3-phenyldicarboxylato-κ5 O, O′:O″,O″,O″′)-bis(N,N-dimethylpropionamide)calcium(II)], C18H25CaN3O8
- Solvothermal synthesis and crystal structure of poly((μ3-2-amino-1,3-benzenedicarboxylato-κ4O:O′,O″,O‴)-bis(μ4-2-amino-1,3-benzenedicarboxylato-κ5O:O:O′,O″,O‴)-(N,N-dimethylformamide)-aqua-di-europium(III)-N,N-dimethylformamide solvate, C30H31N5O7Eu2
- Crystal structure of (1R,4S,6S,6aS,9R,11bS,12S)-1,6,12-trihydroxy-4,11b-dimethyl-8-methylene-7-oxotetradecahydro-6a,9-methanocyclohepta[a]naphthalen-4-yl)methyl acetate
- Crystal structure of diaqua[bis(5-methyl-2-pyrazinecarboxylato-κ2N,O)]zinc(II), C12H14N4O6Zn
- Hydrogen bonding in the crystal structure of the organic salt (R)-1-phenylethan-1-aminium (R)-5-(1,2-dithiolan-3-yl)pentanoate – a salt of the lipoic acid
- The crystal structure of 5-amino-4-(4-amino-1-oxoisoindolin-2-yl)-5-oxopentanoic acid, C13H15N3O4
- Crystal structure of 2-((4-morpholino-1,2,5-thiadiazol-3-yl)oxy)benzaldehyde, C13H13O3N3S
- The crystal structure of 2,8,14,20-tetrapropylpentacyclo[19.3.1.13,7.19,13.115,19]octacosa-1(25),3(28),4,6,9(27),10,12,15(26),16,18, 21,23-dodecaene-4,5,6,10,11,12,16,17,18,22,23,24-dodecol – acetone – water (1(3/3))
- Crystal structure of bis(m2-(2-oxidophenyl)-5-(pyridin-2-yl)pyrazolyl k4N,N′:N′′,O:O)-(μ2-acetic acid k2O:O)-(N,N-dimethylformamide k1O)dimanganese(II) N,N-dimethylformamide solvate, C36H36Mn2N8O6
- The crystal structure of 2,2′-difluoro-3,3′-dimethoxy-1,1′-biphenyl, C14H12F2O2
- The crystal structure of (E)-4-(2-methoxyphenoxy)-3-(2-(-1,3,3,6a-tetramethyl-8-methylenedecahydro-1H-naphtho[2,1-d][1,3]dioxin-7-yl)ethylidene)dihydrofuran-2(3H)-one, C30H40O6
- Crystal structure of 3-hydroxy-2-mesityl-4,4-dimethyl-3,4-dihydroisoquinolin-1(2H)-one – a Z′ = 3 structure –
- Crystal structure of bis(1,1′-biphenyl-2,2′-diolato-κ2O,O′)-bis (1H-pyrazole-κ1N)titanium(IV) pyrazole solvate, C36H32N8O4Ti
- Crystal structure of di-μ-azido -k1N bis[2-(1,10-phenanthrolin-2-yl)benzo[d]imidazol-1-ido-k3N,N′,N′′]-bis(methanol-k1O)-di-manganese
- Crystal structure of bis(acetato-κ1O)-bis methanol-κ1O)bis(1-(4-chlorophenoxy)-3,3-dimethyl-1-(1H-1,2,4-triazol-1-yl)butan-2-ol-κ1N)nickel(II)
- Crystal structure of (E)-(3-(4-fluorophenyl)acryloyl)ferrocene, C19H15FFeO
- The crystal structure of 5,7-dinitro-1,3,5,7-dioxadiazocane
- Crystal structure of 5-(2,5-dimethoxyphenyl)-8-methoxy-2-oxo-2H-chromene-6-carbaldehyde, C19H16O6
- The crystal structure of 2,5-diphenyloxazole–2,5-dihydroxyterephthalic acid (2/1), C19H14NO4
- Crystal structure of [bis(acetonitrile κ1N)-bis(μ2-3,3′-methylenebis(1-methyl-1,3-dihydro-2H-imidazole-2-thione)-κ2S,S)iron(II) diperchlorate], C22H30Cl2FeN10O8S4
- The crystal structure of N-((4-chloro-5-(3-nitrophenyl)-2H-1,2,3-triazol-2-yl)methyl)acetamide, C11H10ClN5O3
- The crystal structure of 2-benzylsuccinic acid, C11H12O6 Z. Kristallogr. - New Cryst. Struct.
- The crystal structure of tert-butyl 3-(1-benzoyl-5-methyl-3-oxo-4-phenethyl-1,2,3,4-tetrahydropyrazin-2-yl)-1H-indole-1-carboxylate, C33H33N3O4
- The crystal structure of 4-dimethylaminoantipyrine hydrochloride, C13H18N3OCl
- Crystal structure of poly[μ2-3-(pyrazin-2-yl)-1,2,4-triazolato-k3N:N′,N′′)(μ2-formato-k2O:O′)copper(II)], C7H5CuN5O2
- Synthesis and crystal structure of 4-(ethoxycarbonyl)phenyl 2-chloro-4-fluorobenzoate, C16H12ClFO4
- Crystal structure of (4-(3,4-dimethylphenyl)-1H-pyrrole-3-carbonyl)ferrocene, C23H21FeNO
- Crystal structure of (4-(2,4-dimethoxyphenyl)-1H-pyrrole-3-carbonyl)ferrocene, C23H21FeNO3
- The crystal structure of bis(1-(2-chloroethyl)pyrrolidin-1-ium) hexachloridotin(IV), C12H22Cl8N2Sn
- Co-crystallization of isonicotinamide with diphenic acid: formation of a supramolecular adduct
- Erratum
- Erratum
Artikel in diesem Heft
- Frontmatter
- New Crystal Structures
- The crystal structure of ( E )-4-chloro-7-(diethylamino)-3-(3-oxo-3-phenylprop-1-en-1-yl)-2 H -chromen-2-one C22H20ClNO3
- Crystal structure of (5S,6E,8E,10E,13R,15S,16Z)-15,5,13,15-tetrahydroxy-16-methyl-2-aza-1(1,3)-benzenacyclononadecaphane-6,8,10,16-tetraen-3-one, C24H31NO5
- The crystal structure of [bis(6-phenylpyridine-2-carboxylato-κ2O,N)-dihydrate-nickel(II)] ethanol solvate
- The crystal structure of poly[(μ3-2-nitro-1,3-phenyldicarboxylato-κ5 O, O′:O″,O″,O″′)-bis(N,N-dimethylpropionamide)calcium(II)], C18H25CaN3O8
- Solvothermal synthesis and crystal structure of poly((μ3-2-amino-1,3-benzenedicarboxylato-κ4O:O′,O″,O‴)-bis(μ4-2-amino-1,3-benzenedicarboxylato-κ5O:O:O′,O″,O‴)-(N,N-dimethylformamide)-aqua-di-europium(III)-N,N-dimethylformamide solvate, C30H31N5O7Eu2
- Crystal structure of (1R,4S,6S,6aS,9R,11bS,12S)-1,6,12-trihydroxy-4,11b-dimethyl-8-methylene-7-oxotetradecahydro-6a,9-methanocyclohepta[a]naphthalen-4-yl)methyl acetate
- Crystal structure of diaqua[bis(5-methyl-2-pyrazinecarboxylato-κ2N,O)]zinc(II), C12H14N4O6Zn
- Hydrogen bonding in the crystal structure of the organic salt (R)-1-phenylethan-1-aminium (R)-5-(1,2-dithiolan-3-yl)pentanoate – a salt of the lipoic acid
- The crystal structure of 5-amino-4-(4-amino-1-oxoisoindolin-2-yl)-5-oxopentanoic acid, C13H15N3O4
- Crystal structure of 2-((4-morpholino-1,2,5-thiadiazol-3-yl)oxy)benzaldehyde, C13H13O3N3S
- The crystal structure of 2,8,14,20-tetrapropylpentacyclo[19.3.1.13,7.19,13.115,19]octacosa-1(25),3(28),4,6,9(27),10,12,15(26),16,18, 21,23-dodecaene-4,5,6,10,11,12,16,17,18,22,23,24-dodecol – acetone – water (1(3/3))
- Crystal structure of bis(m2-(2-oxidophenyl)-5-(pyridin-2-yl)pyrazolyl k4N,N′:N′′,O:O)-(μ2-acetic acid k2O:O)-(N,N-dimethylformamide k1O)dimanganese(II) N,N-dimethylformamide solvate, C36H36Mn2N8O6
- The crystal structure of 2,2′-difluoro-3,3′-dimethoxy-1,1′-biphenyl, C14H12F2O2
- The crystal structure of (E)-4-(2-methoxyphenoxy)-3-(2-(-1,3,3,6a-tetramethyl-8-methylenedecahydro-1H-naphtho[2,1-d][1,3]dioxin-7-yl)ethylidene)dihydrofuran-2(3H)-one, C30H40O6
- Crystal structure of 3-hydroxy-2-mesityl-4,4-dimethyl-3,4-dihydroisoquinolin-1(2H)-one – a Z′ = 3 structure –
- Crystal structure of bis(1,1′-biphenyl-2,2′-diolato-κ2O,O′)-bis (1H-pyrazole-κ1N)titanium(IV) pyrazole solvate, C36H32N8O4Ti
- Crystal structure of di-μ-azido -k1N bis[2-(1,10-phenanthrolin-2-yl)benzo[d]imidazol-1-ido-k3N,N′,N′′]-bis(methanol-k1O)-di-manganese
- Crystal structure of bis(acetato-κ1O)-bis methanol-κ1O)bis(1-(4-chlorophenoxy)-3,3-dimethyl-1-(1H-1,2,4-triazol-1-yl)butan-2-ol-κ1N)nickel(II)
- Crystal structure of (E)-(3-(4-fluorophenyl)acryloyl)ferrocene, C19H15FFeO
- The crystal structure of 5,7-dinitro-1,3,5,7-dioxadiazocane
- Crystal structure of 5-(2,5-dimethoxyphenyl)-8-methoxy-2-oxo-2H-chromene-6-carbaldehyde, C19H16O6
- The crystal structure of 2,5-diphenyloxazole–2,5-dihydroxyterephthalic acid (2/1), C19H14NO4
- Crystal structure of [bis(acetonitrile κ1N)-bis(μ2-3,3′-methylenebis(1-methyl-1,3-dihydro-2H-imidazole-2-thione)-κ2S,S)iron(II) diperchlorate], C22H30Cl2FeN10O8S4
- The crystal structure of N-((4-chloro-5-(3-nitrophenyl)-2H-1,2,3-triazol-2-yl)methyl)acetamide, C11H10ClN5O3
- The crystal structure of 2-benzylsuccinic acid, C11H12O6 Z. Kristallogr. - New Cryst. Struct.
- The crystal structure of tert-butyl 3-(1-benzoyl-5-methyl-3-oxo-4-phenethyl-1,2,3,4-tetrahydropyrazin-2-yl)-1H-indole-1-carboxylate, C33H33N3O4
- The crystal structure of 4-dimethylaminoantipyrine hydrochloride, C13H18N3OCl
- Crystal structure of poly[μ2-3-(pyrazin-2-yl)-1,2,4-triazolato-k3N:N′,N′′)(μ2-formato-k2O:O′)copper(II)], C7H5CuN5O2
- Synthesis and crystal structure of 4-(ethoxycarbonyl)phenyl 2-chloro-4-fluorobenzoate, C16H12ClFO4
- Crystal structure of (4-(3,4-dimethylphenyl)-1H-pyrrole-3-carbonyl)ferrocene, C23H21FeNO
- Crystal structure of (4-(2,4-dimethoxyphenyl)-1H-pyrrole-3-carbonyl)ferrocene, C23H21FeNO3
- The crystal structure of bis(1-(2-chloroethyl)pyrrolidin-1-ium) hexachloridotin(IV), C12H22Cl8N2Sn
- Co-crystallization of isonicotinamide with diphenic acid: formation of a supramolecular adduct
- Erratum
- Erratum

