Abstract
Objectives
The study aims to improve the classification of fetal anatomical planes using Deep Learning (DL) methods to enhance the accuracy of fetal ultrasound interpretation.
Methods
Five Convolutional Neural Network (CNN) architectures, such as VGG16, ResNet50, InceptionV3, DenseNet169, and MobileNetV2, are evaluated on a large-scale, clinically validated dataset of 12,400 ultrasound images from 1,792 patients. Preprocessing methods, including scaling, normalization, label encoding, and augmentation, are applied to the dataset, and the dataset is split into 80 % for training and 20 % for testing. Each model was fine-tuned and evaluated based on its classification accuracy for comparison.
Results
DenseNet169 achieved the highest classification accuracy of 92 % among all the tested models.
Conclusions
The study shows that CNN-based models, particularly DenseNet169, significantly improve diagnostic accuracy in fetal ultrasound interpretation. This advancement reduces error rates and provides support for clinical decision-making in prenatal care.
Introduction
Prenatal ultrasound [1], [2], [3] is an important, simple imaging method that clinicians use widely to diagnose prenatal defects by assessing the fetus’s health, development, and position. Despite its widespread use, the traditional interpretation of ultrasound images remains complex, time-consuming, and dependent on professional expertise. Structural defects impact about one out of every 40 infants, and early detection is typically critical to effective intervention and care [4]. However, the complexity of embryonic anatomical components [5], the variety in picture quality, and the subjectivity of human interpretation all pose significant challenges to reliable and precise diagnosis.
In recent years, deep learning (DL) [6] has demonstrated itself as a significant method in healthcare, particularly in medical image analysis, by offering automated solutions with noteworthy performance in tasks such as disease detection, segmentation, and classification. Among the DL models, Convolutional Neural Networks (CNNs) [7] have shown outstanding success due to their ability to extract hierarchical features from image data, making them ideal candidates for addressing challenges in fetal ultrasound imaging. Existing studies have applied CNNs to classify fetal planes and detect anatomical structures [8], but several continuous difficulties still limit clinical deployment. Here, Rauf et al. [9] have introduced two novel convolutional neural network architectures, namely the 3-residual and the 4-residual-block models, each designed to enhance efficiency compared to ResNet 18 and ResNet 50. But they face challenges for training a deep learning model in an imbalanced dataset and can lead to the extraction of irrelevant information from deeper layers. Sarno et al. [10] evaluates the potential of AI in obstetrics for improving diagnosis but faces challenges such as the need for larger datasets, clinician training, and evidence-based guidelines for clinical integration. Xie et al. [11] verified the feasibility of classifying fetal brains using DL algorithms for the binary classification of normal or abnormal standard fetal ultrasound brain images in axial planes. However, their used dataset was not multicenter, which limited data diversity. Moreover, they focused only on binary classification (normal vs. abnormal) of fetal brain, lacking generalizability to other anatomical planes. These include limited documented datasets, imbalanced class distributions, difficulty in distinguishing visually similar anatomical regions, and the necessity for model interpretability in clinical decision-making.
This study aims to address these challenges by implementing and evaluating five different CNN architectures [12], such as VGG16, ResNet50, InceptionV3, DenseNet169, and MobileNetV2, on a large dataset (FETAL_PLANES_DB) [13] consisting of over 12,400 ultrasound images from 1,792 patients. In contrast to many existing studies, this research focuses on classification accuracy, the clinical scalability, and explainability of the models, providing the findings more applicable to real-world medical use. The study also demonstrates that high-performance DL in fetal ultrasound analysis can be achieved without the need for expensive computing infrastructure, making it more accessible to low-resource healthcare environments. The main objectives of this study are:
To present a reproducible and efficient DL method for classifying fetal ultrasound images across key anatomical planes using a clinically validated dataset.
To evaluate and compare the performance of five innovative CNN architectures like VGG16, ResNet50, InceptionV3, DenseNet169, and MobileNetV2.
To evaluate the clinical applicability and scalability of the proposed models, providing their real-world diagnostic utility, computational feasibility, and possible incorporation into prenatal care processes.
Several studies have used DL models to enhance prenatal ultrasound image interpretation, focusing on fetal plane categorization, anomaly diagnosis, and structure segmentation. This section reviews some existing methods, findings, and challenges. Rathika et al. [14] present a Radial Basis Function Neural Network (RBFNN) method and achieves a classification accuracy of 98.07 %. The study needs high computational resources and doesn’t explore deployment for real-time clinical applications. Yang et al. [15] collected 1779 normal and abnormal fetal US cardiac pictures in five standard views of the heart. They employed five You Only Look Once version 5 (YOLOv5) networks as their primary model to categorize photos as “normal” or “abnormal”. According to the study, their model achieved an overall accuracy of 90.67 %. Gong et al. [16] developed a novel generative adversarial network (GAN) model by integrating deep anomaly detection (DANomaly) and generative adversarial CNN (GACNN) architectures to detect fetal congenital heart disease (FCH) from echocardiography images. Using a modified WGAN-GP, they created the DGACNN model, achieving 85 % accuracy in detecting FCH.
Zhou et al. [17] introduced the Category Attention-Instance Segmentation Network for segmenting fetal cardiac four-chamber ultrasound images. By adding a Category Attention Module, the model reduces misclassification errors. However, they faced challenges in distinguishing individual instances and needed better context modeling on a more diverse dataset. Nurmaini et al. [18] analyzed four common heart images and three congenital heart abnormalities while utilizing a Mask-RCNN architecture to predict 24 features in fetal heart sectors. The model achieved a DICE score of 89.70 % and an IoU of 79.97 %. However, it was tested on a small dataset of 1,149 fetal heart images. Moradi et al. [19] proposed Multi-Feature Pyramid U-net (MFP-Unet) for automated left ventricle segmentation in 2D echocardiography. Trained on an augmented dataset of 1,370 images, MFP-Unet achieved a Dice score of 0.945 and Hausdorff Distance of 1.62. However, the study was limited by dataset size, image resolution, and computational resources. Qu et al. [20] introduced a differential CNN method for identifying six fetal brain planes by computing the element-wise difference between input images. Although they achieved 92.93 % accuracy using data augmentation, the approach is sensitive to equipment and human operational variations, which can affect model robustness and diagnostic precision. Xie et al. [21] developed a model using DCNN, U-Nets, and VGG-Net to identify embryonic brain abnormalities in fetal images, achieving an overall accuracy of 91.5 % and high F1-scores. Class activation mapping (CAM) helped visualize effectively, but the low IoU values indicate that better object detection methods are needed for more accurate localization.
Wang et al. [22] developed FB-ZWUNet to improve the cerebellum segmentation in fetal brain ultrasounds, achieving a Dice coefficient of 0.8743 and IoU of 0.7813. However, its performance on different ultrasound devices and poor-quality images remains untested. Ciobanu et al. [23] developed an automated method for classifying fetal abdominal planes using MobileNetV3Large and EfficientNetV2S, with accuracies of 79.89 % and 79.19 %. Table 1 presents a comparison of the existing methods and their respective performance metrics.
Existing fetal anomaly detection methods and their performance metrics.
| Reference | Dataset | Features extraction | Model | Results | Limitations |
|---|---|---|---|---|---|
| Rathika et al. [14] | Fetal dataset from Zenodo | Gray-level Co-occurrence matrix | RBFNN | Accuracy: 98.07 % | Higher computational resources, no clinical relevance |
| Yang et al. [15] | Normal and abnormal fetal ultrasound heart images | YOLOv5 component outputs | YOLOv5 models | Accuracy: 90.67 % | Small sample size, class imbalace |
| Gong et al. [16] | Fetal cardiac ultrasound dataset | Region of interest, end-systolic phase | DGACNN | Accuracy: 85 % | Limited disease samples, unlabeled video data |
| Zhou et al. [17] | 319 fetal echocardiography images. | CA-ISNet | Category attention module | Precision: 45.64 %, DICE: 0.7470 to 0.8199 | SOLOv2 misclassification Issue.Moderate precision |
| Nurmaini et al. [18] | Ultrasound video data (1,149 annotated images) | Deep visual and semantic features extracted via ResNet50 | Mask RCNN architecture | DICE=89.70 % and IoU=79.97 | Small dataset |
| Moradi et al. [19] | Clinical images of 500 patients | Grayscale and contrast-enhanced channels. | MFP-Unet | DM: 0.953, HD: 3.49, MAD: 1.12 | Dataset size and Diversity, Limited input image resolution |
| Qu et al. [20] | Fetal brain ultrasound images | Differential convolutional feature | Differential CNN | Accuracy: 92.93 % | dataset’s size and diversity, sensitive to inherent artifacts |
| Xie et al. [21] | Ultrasound fetal images | Extracted spatial features (edges, textures, contours) | U-Net, VGG-Net-based DCNN model | Overall accuracy: 91.5 % | Localization Precision, considered transverse standard planes, neglecting sagittal and coronal planes |
| Wang et al. [22] | Fetal brain corpus callosum dataset | ZAM,WAM, MCM | FB-ZWUNet | Dice: 0.8743, IoU: 0.7813 | Device-specific performance, Limited generalizability |
| Ciobanu et al. [23] | Prospective cohort of fetal ultrasound images (9 classes) | Optical character recognition | MobileNetV3 large, efficient NetV2S | MobileNetV3 large: 79.89 %, EfficientV2S: 79.19 % | Abdominal-only focus, no real-time integration |
Methods
The study addresses the challenging task of classifying ultrasound images into five distinct anatomical categories. Figure 1 illustrates the entire process, from data collection and preprocessing to model training and performance evaluation. A detailed step-by-step process for fetal ultrasound image processing – covering data collection, model training, evaluation, and prediction – is presented in Algorithm 1 (Table 2).
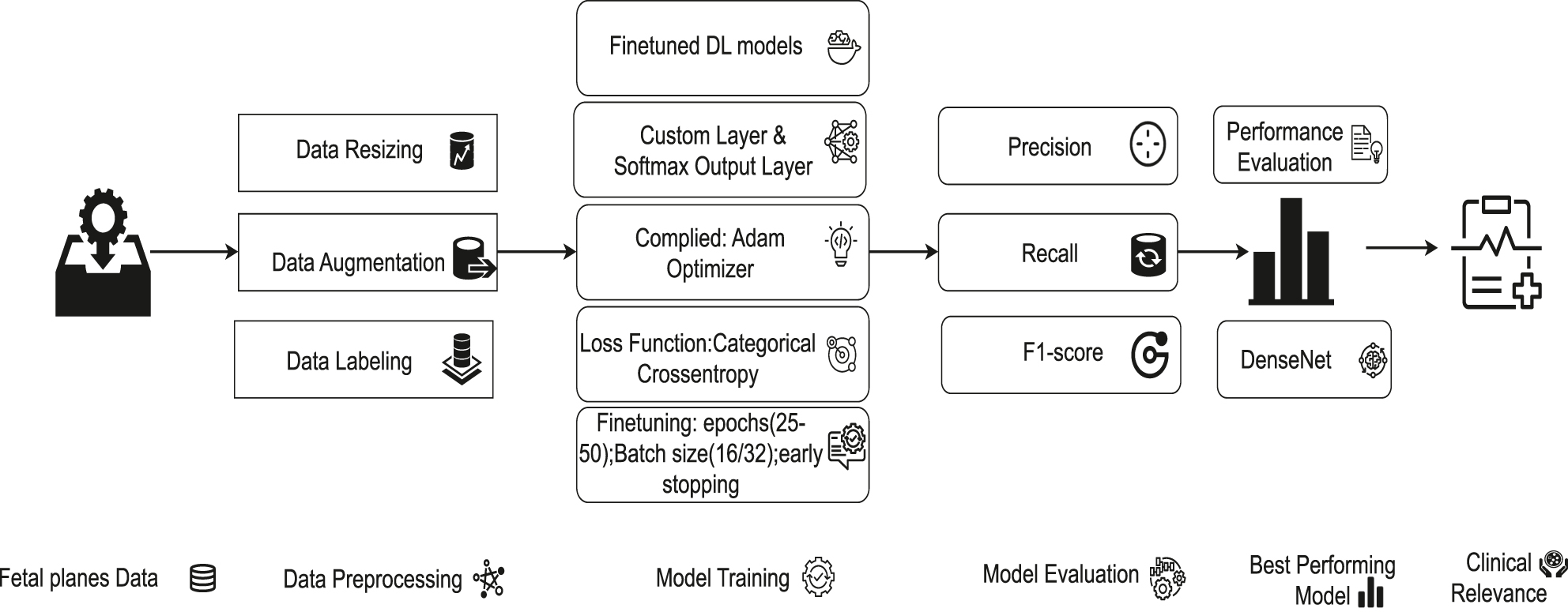
Architectural diagram of fetal ultrasound classification.
Algorithm 1: Working procedure for classification of fetal ultrasound images using DL models.
| 1: Input: Raw ultrasound images from the FETAL_PLANES_DB dataset |
| 2: Step 1: Data Collection |
| 3: Acquire ultrasound images labeled into six categories: Fetal abdomen, fetal brain, fetal femur, fetal thorax, maternal cervix, and other. |
| 4: Step 2: Data Preprocessing |
| 5: Resize all images to fixed dimensions (150 × 150 or 224 × 224 based on model type). |
| 6: Convert grayscale images to 3-channel RGB format. |
| 7: Normalize pixel values to the range [−1, 1]. |
| 8: Apply augmentation (rotation, flipping, zooming, shearing) to increase variability. |
| 9: Encode labels using LabelEncoder and one-hot encoding. |
| 10: Step 3: Dataset Splitting |
| 11: Divide data into training (80 %) and testing (20 %) sets. |
| 12: Step 4: Model Selection and Customization |
| 13: Utilize customized CNN models (e.g., VGG16, ResNet50, InceptionV3, DenseNet169, MobileNetV2). |
| 14: Remove original top layers and add new Dense + Dropout + Softmax layers. |
| 15: Step 5: Model Compilation |
| 16: Use Adam optimizer and categorical cross-entropy loss. |
| 17: Set evaluation metric as classification accuracy. |
| 18: Step 6: Training and Fine-Tuning |
| 19: Train each model for 25–50 epochs with batch size of 16 or 32. |
| 20: Apply early stopping and learning rate scheduling. |
| 21: Step 7: Evaluation |
| 22: Evaluate trained models using accuracy, precision, recall, F1-score, and confusion matrix. |
| 23: Identify best-performing architecture (DenseNet169) based on evaluation metrics. |
| 24: Return: Predicted anatomical category for each input image. |
Data collection
This study utilizes the (FETAL_PLANES_DB) [13] consists of 1,792 patient records with 12,000+ ultrasound images in a real clinical environment. The six classes represent clinically significant fetal structures: (i) fetal abdomen, (ii) fetal brain, (iii) fetal femur, (iv) fetal thorax, (v) maternal cervix, and (vi) Other. The distribution of each type of fetal ultrasound image is as follows: fetal abdomen: 5.7 %, fetal brain: 24.9 %, maternal cervix: 13.1 %, fetal femur: 8.4 %, fetal thorax: 13.9 %, Other: 33.9 %. Figure 2 shows the different maternal–fetal anatomical planes [24] and data representation before and after preprocessing. Each image was labelled by an Expert clinician to ensure high-quality annotations. The images in the dataset vary in size, which can lead to inconsistent visual representation and negatively impact the quality of analysis and model performance. Therefore, resizing them to a uniform dimension is essential for maintaining clarity and consistency.

Visual representation of six different fetal ultrasound images before and after data preprocessing. (A) Different maternal–fetal anatomical planes and data representation before preprocessing, (B) different maternal–fetal anatomical planes after preprocessing.
Data preprocessing
In the preprocessing stages, images are resized using OpenCV to 150 × 150 pixels for models like VGG16 and ResNet50 and to 224 × 224 pixels for models such as InceptionV3, DenseNet169, and MobileNetV2. The grayscale ultrasound image is converted to RGB by repeating the single channel three times for transfer learning with ImageNet-trained weights. Then the images are normalized from 8 bit pixel values [0, 255] to the range [−1, 1] for faster convergence and to improve the training process. The final step in preprocessing is data augmentation, including random rotation, zooming, shearing, and horizontal and vertical flipping, which is used only on the training set to increase model generalization and reduce overfitting. Then, the categorical text labels are first transformed to integer representations using Scikit-Learn’s LabelEncoder function. This stage converts each class into a distinct numerical value. These integer-encoded labels are then turned into a single hot-encoded vector using the to_categorical() function in TensorFlow or Keras [25], which is required for multi-class classification using softmax-based [26] neural network [25] outputs.
Model training
This study optimized the models to separate fetal ultrasound images into five anatomical groups. Each model has been configured with ImageNet [27] trained weights, enabling transfer learning [27] by utilizing pretrained visual features. The final layers of each model are customized to the current classification task by replacing the original fully connected layers with new dense layers, followed by a softmax activation function for multi-class prediction. The Adam optimizer [28] is chosen here due to its adjustable learning rate and computing efficiency, which make it most effective in medical imaging tasks with a large number of parameters. Each model is trained over 25 to 50 epochs with a batch size of 16 or 32, based on model size, early stopping criteria, and memory usage. Figure 3 shows the structured model training process of this study.

Comprehensive architecture of model training process with parameter specifications.
VGG16
The VGG16 [12] architecture is initialized using weights obtained from training on the ImageNet dataset and configured without its top classification layer. Equation (1) represents the fundamental convolutional operation used in the VGG16 architecture. Here, Z(l) represent the output of the lth layer in the network. The terms W(l) and b(l) denote the weights and biases associated with that layer, respectively. The symbol ∗ refers to the convolution operation applied between the input and the kernel. The activation function used is the Rectified Linear Unit (ReLU), mathematically defined as ReLU(x) = max(0, x), which introduces non-linearity into the model while maintaining computational efficiency. It uses small 3 × 3 kernels to capture fine-grained spatial characteristics and recognize features such as the head, abdomen, or femur to distinguish fetal planes.
ResNet50
ResNet50 [12] is initialized with trained weights of ImageNet and modified by appending a custom classification head. Equation (2) represents the fundamental operation used in the ResNet50 architecture. Here, Z(l) is the input to the residual block, and Z(l+1) is the output. The function F(Z(l), W(l)) represents the residual mapping learned by the block, typically consisting of a series of convolutional layers with ReLU activations and batch normalization. The original input Z(l) is added directly to the output of F, enabling the network to learn modifications to the identity function. It is used to preserve important details in deep layers, effectively detect small differences in fetal images, and accurately distinguish between similar fetal planes.
InceptionV3
InceptionV3 [12] model is initialized with trained ImageNet weights, and its top layers are excluded to allow the addition of a task-specific classification head. Equation (3) represents the fundamental operation used in the DenseNet169 architecture. Here, X represents the input, Convk×k denote convolution with a k × k kernel, Pool3×3 represents 3 × 3 max or average pooling, and Concat indicates depth-wise concatenation of feature maps. The model improves performance by using a learning rate scheduler, early stopping, model checkpointing, Adam optimization, and categorical cross-entropy.
InceptionV3 incorporates multiple responses in parallel, making it highly effective for capturing features at different levels. This is particularly relevant in fetal ultrasound where organs and anatomical regions vary significantly in size and orientation. Its efficiency and robustness help in learning discriminative features across varying fetal positions and scanning angles.
DenseNet169
The DenseNet169 [12] model is initialized with ImageNet pre-trained weights and configured without its top classification layers. Eq. (4) represents the fundamental operation used in the DenseNet169 architecture. Here, H(l) is a composite function (BatchNorm → ReLU → Convolutionlayer) and [Z(0), Z(1), …, Z(l−1)] represents concatenation of all previous layer outputs. DenseNet169 improves feature reuse and gradient to detect subtle anatomical differences in fetal ultrasound images.
MobileNetV2
The MobileNetV2 [12] model is trained using a transfer learning approach, where the base convolutional layers are initialized with pre-trained weights from the ImageNet dataset and all layers are halted to allow fine-tuning. Equation (5) represents the fundamental operation used in the MobileNetV2 architecture. Here, ConvDepthwise(X) applies spatial filtering per channel, ConvPointwise performs 1 × 1 convolution for cross-channel fusion, and ConvDS(X) represents the output.
Results
All experiments are conducted on a computational system equipped with an Intel(R) Core(TM) i5-10400 CPU @ 2.90 GHz, 16 GB DDR4 RAM, and a 64 bit Windows 11 operating system (×64-based architecture), ensuring reproducibility and alignment with standard hardware configurations for model evaluation. The model’s performance [29] is evaluated using accuracy, precision, recall, F1-score, and confusion matrices. Each metric is computed based on the number of correctly and incorrectly predicted samples, categorized as True Positives (TP), True Negatives (TN), False Positives (FP), and False Negatives (FN). This ensured a thorough and reliable assessment of its classification effectiveness. Equation (6)–(9) shows the calculating procedure of the evaluation metrics.
Performance metrics
The VGG16 model, fine-tuned with pre-trained ImageNet weights, achieved a test accuracy of 90.22 %. The ResNet50 model, with a test accuracy of 88 %, is close to VGG16, while InceptionV3 also performs similarly with an accuracy close to 90 %. MobileNetV2, optimized for speed and efficiency, achieved a 71 % accuracy. Among all models, DenseNet169 outperforms the others with the highest test accuracy of 92 % presented in Table 3.
Comparison of classification reports for different deep learning models.
| VGG16 classification report | ||||
|---|---|---|---|---|
| Class | Precision | Recall | F1-score | Support |
| Fetal abdomen | 0.77 | 0.72 | 0.74 | 154 |
| Fetal brain | 0.97 | 0.97 | 0.97 | 625 |
| Fetal femur | 0.85 | 0.84 | 0.84 | 199 |
| Fetal thorax | 0.83 | 0.85 | 0.84 | 319 |
| Maternal cervix | 0.99 | 0.98 | 0.98 | 339 |
| Other | 0.88 | 0.89 | 0.88 | 828 |
|
|
||||
| ResNet50 classification report | ||||
|
|
||||
| Class | Precision | Recall | F1-score | Support |
|
|
||||
| Fetal abdomen | 0.66 | 0.60 | 0.63 | 154 |
| Fetal brain | 0.96 | 0.96 | 0.96 | 625 |
| Fetal femur | 0.83 | 0.80 | 0.82 | 199 |
| Fetal thorax | 0.77 | 0.87 | 0.82 | 319 |
| Maternal cervix | 0.98 | 0.99 | 0.98 | 339 |
| Other | 0.88 | 0.86 | 0.87 | 828 |
|
|
||||
| InceptionV3 classification report | ||||
|
|
||||
| Class | Precision | Recall | F1-score | Support |
|
|
||||
| Fetal abdomen | 0.76 | 0.77 | 0.76 | 154 |
| Fetal brain | 0.98 | 0.97 | 0.98 | 625 |
| Fetal femur | 0.82 | 0.80 | 0.81 | 199 |
| Fetal thorax | 0.85 | 0.83 | 0.84 | 319 |
| Maternal cervix | 1.00 | 0.99 | 0.99 | 339 |
| Other | 0.88 | 0.89 | 0.89 | 828 |
|
|
||||
| MobileNetV2 classification report | ||||
|
|
||||
| Class | Precision | Recall | F1-score | Support |
|
|
||||
| Fetal abdomen | 1.00 | 0.41 | 0.58 | 142 |
| Fetal brain | 0.71 | 1.00 | 0.83 | 615 |
| Fetal femur | 0.62 | 0.20 | 0.31 | 207 |
| Fetal thorax | 1.00 | 0.45 | 0.62 | 274 |
| Maternal cervix | 0.43 | 1.00 | 0.60 | 322 |
| Other | 1.00 | 0.64 | 0.78 | 839 |
|
|
||||
| DenseNet169 classification report | ||||
|
|
||||
| Class | Precision | Recall | F1-score | Support |
|
|
||||
| Fetal abdomen | 0.79 | 0.84 | 0.81 | 154 |
| Fetal brain | 0.97 | 0.99 | 0.98 | 625 |
| Fetal femur | 0.88 | 0.77 | 0.82 | 199 |
| Fetal thorax | 0.89 | 0.89 | 0.89 | 319 |
| Maternal cervix | 0.99 | 1.00 | 0.99 | 339 |
| Other | 0.89 | 0.89 | 0.89 | 828 |
Figure 4 shows the confusion matrix of all five CNN models. Figure 5 shows high classification accuracy for all classes, with AUC values between 0.87 and 1.00.

Confusion matrix of various CNN models’. (A) VGG16 model performance. (B) ResNet50 model performance. (C) InceptionV3 model performance. (D) MobileNetV2 model performance. (E) DenseNet169 model performance.

ROC curves of various CNN architectures used for fetal anatomical plane classification. (A) VGG16 model ROC curve. (B) ResNet50 model ROC curve. (C) InceptionV3 model ROC curve. (D) MobileNetV2 model ROC curve. (E) DenseNet169 model ROC curve.
Comparative analysis
We compared the classification performance of various CNN architectures on fetal ultrasound image data, using the models VGG16, ResNet50, InceptionV3, DenseNet169, and MobileNetV2. Table 4 presents the performance metrics, which include overall accuracy, average precision, recall, F1-score, and best performing classes. DenseNet169 outperformed all other models as a result of accuracy and F1-score, demonstrating an outstanding capacity to identify complex anatomical structures like the fetal thorax. In comparison, MobileNetV2, performed considerably lower in most metrics, including detecting fetal abdomen and femur. The findings demonstrate the relationship between model complexity and diagnostic performance.
Performance comparison of CNN models for ultrasound classification.
| Model | Accuracy, % | Precision, P | Recall, R | F1-score | Best performing classes | Weakest classes | Computational efficiency |
|---|---|---|---|---|---|---|---|
| VGG16 | 90.22 | 0.88 | 0.88 | 0.88 | Fetal brain, maternal cervix | Fetal abdomen (77 % P, 72 % R), fetal thorax | Moderate |
| ResNet50 | 88.00 | 0.86 | 0.86 | 0.86 | Fetal brain, maternal cervix | Fetal abdomen (66 % P, 60 % R) | Moderate |
| InceptionV3 | 90.00 | 0.89 | 0.89 | 0.89 | Fetal brain, maternal cervix | Fetal abdomen (76 % P, 77 % R), fetal thorax | High |
| DenseNet169 | 92.00 | 0.90 | 0.90 | 0.90 | Fetal brain, maternal cervix, fetal thorax | Fetal abdomen (79 % P, 84 % R) | High |
| MobileNetV2 | 65.83 | 0.79 | 0.62 | 0.62 | Fetal brain | Fetal abdomen (41 % P, 41 % R), fetal thorax, fetal femur | Lightweight |
Discussions
This study conducts a comprehensive evaluation of five deep learning models, VGG16, ResNet50, InceptionV3, DenseNet169, and MobileNetV2 for the classification of fetal ultrasound images across six anatomical planes. Among these models, DenseNet169 demonstrated superior performance, achieving a classification accuracy of 92 %, thereby highlighting its exceptional capability in recognizing intricate fetal structures, particularly the fetal brain and maternal cervix. InceptionV3 and VGG16 also exhibited competitive performance. In contrast, MobileNetV2, despite its lightweight and computationally efficient design, significantly underperformed with an accuracy of 65.83 %, especially in complex anatomical regions such as the fetal abdomen and femur. This study identified deeper CNN architectures, particularly DenseNet169, outperforming lighter models such as MobileNetV2 in classifying complex fetal ultrasound images. This indicates deeper networks are more effective at preserving subtle spatial features and fine-grained anatomical distinctions, which are important for accurate classification in medical imaging tasks. These findings are important because they demonstrate the clinical feasibility of incorporating advanced deep learning models into diagnostic workflows. DenseNet169, with high F1-scores across most classes, may be utilized as an option for sonographers, lowering diagnostic errors, particularly in resource-constrained environments. MobileNetV2’s low performance, despite being fast and efficient, shows the limitations of using lightweight models in high-precision tasks without customized optimizations. The models also show that almost all models performed better when recognizing the fetal brain and maternal cervix, suggesting that these anatomical planes are visually separate and simpler to segment. On the other hand, the fetal abdomen consistently appeared as the weakest class across all models, with significantly lower precision and recall. The inability of models to learn robust features is probably caused by both class imbalance and increased variation between different classes. The DenseNet169 model’s strong performance is significant not just for accuracy but also for usage in real-world medical contexts. Using this type of DL model in prenatal care can help in various ways, including reducing diagnostic time by automatically detecting important anatomical views, improving consistency by reducing differences between different doctors’ interpretations, and assisting doctors in making correct decisions. These advantages are particularly useful in clinics with limited resources, where professional doctors may not always be available.
Furthermore, while some studies have examined prenatal defects, authors frequently focused on specific structures; this research has utilized a complete multi-class categorization method. Compared to previous studies that primarily addressed limited anatomy or binary classification tasks, this study takes a more comprehensive approach. Our methodology covers multi-class categorization across six essential fetal anatomical planes, hence increasing its therapeutic value in real-world circumstances. Furthermore, previous research frequently did not include robust testing across varied anatomical complexity and failed to incorporate multi-plane detection. Our findings highlight the relevance of employing deeper CNN designs such as DenseNet169, which remains successful across almost all classes. This contributes to broader diagnostic applicability and sets a benchmark for automated fetal ultrasound classification systems.
Although the dataset is clinically labeled and large, it may not cover every aspect of fetal morphological changes among populations. This limitation could affect the model’s generalization when applied to new clinical settings. The training and evaluation were conducted on resource-constrained hardware. While this demonstrates the feasibility of Deep Learning on modest systems, it may have limited the exploration of larger architectures or more complex hyperparameter tuning. A slight imbalance in the number of samples across classes, particularly for the fetal abdomen and fetal femur, may have influenced the model’s performance. Models may have been biased toward classes with more examples, resulting in reduced recall and precision in underrepresented categories. To extend the scope and applicability of this study, future research will focus on several key areas. Firstly, expanding the dataset with fetal ultrasound images from multiple hospitals, different ultrasound machines, and varied patient populations will enhance the model’s generalization and robustness across diverse clinical settings. Secondly, exploring model compression and optimization strategies such as pruning, quantization, or the use of inherently lightweight architectures like MobileNetV2 can facilitate real-time deployment in environments with limited computational resources. Third, improving the explainability of model decisions through visualization techniques like Grad-CAM or SHAP can increase transparency and help clinicians understand and trust the model’s outputs. Lastly, integrating these models into real-time clinical workflows should be investigated. Real-time deployment during ultrasound scans has the potential to support practitioners with immediate feedback, thus improving both the speed and consistency of prenatal diagnostics.
Conclusions
This study proposes a deep learning based method for the classification of fetal ultrasound images using five customized CNN architectures. These models are trained and evaluated on the FETAL_PLANES_DB dataset, which contains six clinically significant anatomical planes. The objective is to evaluate the effectiveness of current deep learning approaches in accurately identifying fetal structures from ultrasound images. Among the models, DenseNet169 achieves the highest test accuracy of 92 %. The results show the need for deep learning models to support automated prenatal diagnostics, enabling clinicians to identify fetal anatomical features more efficiently and with reduced variability. This method performs effectively even on basic resource equipment, demonstrating its suitability for deployment in low-resource clinical environments. To provide higher reliability in real-world applications, future research should focus on increasing dataset diversity, optimizing model performance, and improving interpretability. Improving these fields is essential for incorporating deep models into reliable and efficient medical procedures.
-
Research ethics: Not applicable.
-
Informed consent: Not applicable.
-
Author contributions: All authors have accepted responsibility for the entire content of this manuscript and approved its submission.
-
Use of Large Language Models, AI and Machine Learning Tools: None declared.
-
Conflict of interest: The authors state no conflict of interest.
-
Research funding: None declared.
-
Data availability: The datasets analyzed during the current study are available in the ZENODO repository, https://zenodo.org/records/3904280.
References
1. He, F, Wang, Y, Xiu, Y, Zhang, Y, Chen, L. Artificial intelligence in prenatal ultrasound diagnosis. Front Med 2021;8. https://doi.org/10.3389/fmed.2021.729978.Search in Google Scholar PubMed PubMed Central
2. Rizzo, G, Pietrolucci, ME, Capponi, A, Mappa, I. Exploring the role of artificial intelligence in the study of fetal heart. Int J Cardiovasc Imag 2022;38:1017–9. https://doi.org/10.1007/s10554-022-02588-x.Search in Google Scholar PubMed
3. Pietrolucci, ME, Maqina, P, Mappa, I, Marra, MC, D’Antonio, F, Rizzo, G. Evaluation of an artificial intelligent algorithm (Heartassist™) to automatically assess the quality of second trimester cardiac views: a prospective study. J Perinat Med 2023;51:920–4. https://doi.org/10.1515/jpm-2023-0052.Search in Google Scholar PubMed
4. Dey, SK, Uddin, KM, Howlader, A, Rahman, MM, Babu, HM, Biswas, N, et al.. Analyzing infant cry to detect birth asphyxia using a hybrid CNN and feature extraction approach. Neurosci Inf 2025;5.10.1016/j.neuri.2025.100193Search in Google Scholar
5. Hikspoors, JP, Kruepunga, N, Mommen, GM, Köhler, SE, Anderson, RH, Lamers, WH. A pictorial account of the human embryonic heart between 3.5 and 8 weeks of development. Commun Biol 2022;5:226. https://doi.org/10.1038/s42003-022-03153-x.Search in Google Scholar PubMed PubMed Central
6. Ramirez Zegarra, R, Ghi, T. Use of artificial intelligence and deep learning in fetal ultrasound imaging. Ultrasound Obstet Gynecol 2023;62:185–94. https://doi.org/10.1002/uog.26130.Search in Google Scholar PubMed
7. Liang, H, Lu, Y. A CNN-RNN unified framework for intrapartum cardiotocograph classification. Comput Methods Progr Biomed 2023;229. https://doi.org/10.1016/j.cmpb.2022.107300.Search in Google Scholar PubMed
8. Zhang, Y, Xie, R, Beheshti, I, Liu, X, Zheng, G, Wang, Y, et al.. Improving brain age prediction with anatomical feature attention-enhanced 3D-CNN. Comput Biol Med 2024;169. https://doi.org/10.1016/j.compbiomed.2023.107873.Search in Google Scholar PubMed
9. Rauf, F, Attique Khan, M, Albarakati, HM, Jabeen, K, Alsenan, S, Hamza, A, et al.. Artificial intelligence assisted common maternal fetal planes prediction from ultrasound images based on information fusion of customized convolutional neural networks. Front Med 2024;11. https://doi.org/10.3389/fmed.2024.1486995.Search in Google Scholar PubMed PubMed Central
10. Sarno, L, Neola, D, Carbone, L, Saccone, G, Carlea, A, Miceli, M, et al.. Use of artificial intelligence in obstetrics: not quite ready for prime time. Am J Obstet Gynecol MFM 2023;5. https://doi.org/10.1016/j.ajogmf.2022.100792.Search in Google Scholar PubMed
11. Xie, H, Wang, N, He, M, Zhang, L, Cai, H, Xian, J, et al.. Using deep-learning algorithms to classify fetal brain ultrasound images as normal or abnormal. Ultrasound Obstet Gynecol 2020;56:579–87. https://doi.org/10.1002/uog.21967.Search in Google Scholar PubMed
12. Uddin, KM, Dey, SK, Babu, HM, Mostafiz, R, Uddin, S, Shoombuatong, W, et al.. Feature fusion based VGGFusionNet model to detect COVID-19 patients utilizing computed tomography scan images. Sci Rep 2022;12. https://doi.org/10.1038/s41598-022-25539-x.Search in Google Scholar PubMed PubMed Central
13. Burgos-Artizzu, XP, Coronado-Gutierrez, D, Valenzuela-Alcaraz, B, Bonet-Carne, E, Eixarch, E, Crispi, F, et al.. FETAL_PLANES_DB: common maternal-fetal ultrasound images. Barcelona, Spain: Zenodo; 2020.Search in Google Scholar
14. Rathika, S, Mahendran, K, Sudarsan, H, Ananth, SV. Novel neural network classification of maternal fetal ultrasound planes through optimized feature selection. BMC Med Imag 2024;24:337. https://doi.org/10.1186/s12880-024-01453-8.Search in Google Scholar PubMed PubMed Central
15. Yang, Y, Wu, B, Wu, H, Xu, W, Lyu, G, Liu, P, et al.. Classification of normal and abnormal fetal heart ultrasound images and identification of ventricular septal defects based on deep learning. J Perinat Med 2023;51:1052–8. https://doi.org/10.1515/jpm-2023-0041.Search in Google Scholar PubMed
16. Gong, Y, Zhang, Y, Zhu, H, Lv, J, Cheng, Q, Zhang, H, et al.. Fetal congenital heart disease echocardiogram screening based on DGACNN: adversarial one-class classification combined with video transfer learning. IEEE Trans Med Imag 2019;39:1206–22. https://doi.org/10.1109/tmi.2019.2946059.Search in Google Scholar
17. An, S, Zhu, H, Wang, Y, Zhou, F, Zhou, X, Yang, X, et al.. A category attention instance segmentation network for four cardiac chambers segmentation in fetal echocardiography. Comput Med Imag Graph 2021;93. https://doi.org/10.1016/j.compmedimag.2021.101983.Search in Google Scholar PubMed
18. Nurmaini, S, Rachmatullah, MN, Sapitri, AI, Darmawahyuni, A, Tutuko, B, Firdaus, F, et al.. Deep learning-based computer-aided fetal echocardiography: application to heart standard view segmentation for congenital heart defects detection. Sensors 2021;21:8007. https://doi.org/10.3390/s21238007.Search in Google Scholar PubMed PubMed Central
19. Moradi, S, Oghli, MG, Alizadehasl, A, Shiri, I, Oveisi, N, Oveisi, M, et al.. MFP-Unet: a novel deep learning based approach for left ventricle segmentation in echocardiography. Phys Med 2019;67:58–69. https://doi.org/10.1016/j.ejmp.2019.10.001.Search in Google Scholar PubMed
20. Qu, R, Xu, G, Ding, C, Jia, W, Sun, M. Standard plane identification in fetal brain ultrasound scans using a differential convolutional neural network. IEEE Access 2020;8:83821–30. https://doi.org/10.1109/access.2020.2991845.Search in Google Scholar
21. Xie, B, Lei, T, Wang, N, Cai, H, Xian, J, He, M, et al.. Computer-aided diagnosis for fetal brain ultrasound images using deep convolutional neural networks. Int J Comput Assist Radiol Surg 2020;15:1303–12. https://doi.org/10.1007/s11548-020-02182-3.Search in Google Scholar PubMed
22. Wang, Q, Zhao, D, Ma, H, Liu, B. FB-ZWUNet: a deep learning network for corpus callosum segmentation in fetal brain ultrasound images for prenatal diagnostics. Biomed Signal Process Control 2025;104. https://doi.org/10.1016/j.bspc.2025.107499.Search in Google Scholar
23. Ciobanu, ŞG, Enache, IA, Iovoaica-Rămescu, C, Berbecaru, EIA, Vochin, A, Băluţă, ID, et al.. Automatic identification of fetal abdominal planes from ultrasound images based on deep learning. J Imaging Inf Med 2025:1–8.10.1007/s10278-025-01409-6Search in Google Scholar PubMed
24. Burgos-Artizzu, XP, Coronado-Gutiérrez, D, Valenzuela-Alcaraz, B, Bonet-Carne, E, Eixarch, E, Crispi, F, et al.. Evaluation of deep convolutional neural networks for automatic classification of common maternal fetal ultrasound planes. Sci Rep 2020;10.10.1038/s41598-020-67076-5Search in Google Scholar PubMed PubMed Central
25. Grattarola, D, Alippi, C. Graph neural networks in tensorflow and keras with spektral [application notes]. IEEE Comput Intell Mag 2021;16:99–106. https://doi.org/10.1109/mci.2020.3039072.Search in Google Scholar
26. Selem, M, Jemili, F, Korbaa, O. Deep learning for intrusion detection in IoT networks. Peer-to-Peer Netw Appl 2025;18:22. https://doi.org/10.1007/s12083-024-01819-3.Search in Google Scholar
27. Morid, MA, Borjali, A, Del Fiol, G. A scoping review of transfer learning research on medical image analysis using ImageNet. Comput Biol Med 2021;128. https://doi.org/10.1016/j.compbiomed.2020.104115.Search in Google Scholar PubMed
28. Ogundokun, RO, Maskeliunas, R, Misra, S, Damaševičius, R. Improved CNN based on batch normalization and adam optimizer. In: International Conference on Computational Science and Its Applications. Springer; 2022:593–604 pp.10.1007/978-3-031-10548-7_43Search in Google Scholar
29. Taifa, IA, Setu, DM, Islam, T, Dey, SK, Rahman, T. A hybrid approach with customized machine learning classifiers and multiple feature extractors for enhancing diabetic retinopathy detection. Healthc Anal 2024;5. https://doi.org/10.1016/j.health.2024.100346.Search in Google Scholar
© 2025 the author(s), published by De Gruyter, Berlin/Boston
This work is licensed under the Creative Commons Attribution 4.0 International License.
Articles in the same Issue
- Frontmatter
- Reviews
- Fetal neurobehavior and consciousness: a systematic review of 4D ultrasound evidence and ethical challenges
- Prenatal maternal stress and long-term neurodevelopmental outcomes: a narrative review
- From exencephaly to anencephaly: a catastrophic continuum of neural tube defects from embryogenesis to ultrasonographic diagnosis
- Opinion Papers
- Fetoception: a window into maternal interoception?
- Rationale for the use of fetal ventriculosubgaleal shunts for the treatment of aqueduct stenosis
- Original Articles – Obstetrics
- The fetal occiput-spine angle measurement during first stage of labor as a predictor for vaginal delivery, a systematic review and meta-analysis
- Hemorrhage-related maternal morbidity of secondary compared to primary postpartum hemorrhage
- First-trimester maternal serum PAPP-A levels and hyperemesis gravidarum: unraveling the link – a meta-analysis
- Antepartum cerebroplacental ratio in low risk pregnancy and its relationship with adverse perinatal outcome: a prospective cohort study
- Liver fibrosis markers as predictors of adverse outcomes in pregnancy-related hypertensive disorders
- Endocrine disrupting chemicals: translating mechanisms into perinatal risk assessment
- Low uterine segment thickness in the prediction of cesarean delivery after induction of labor
- Original Articles – Fetus
- Fetal adrenal gland size in preeclamptic pregnancies with and without fetal growth restriction: an ultrasonographic assessment
- Evaluation of safety and performance of CentaFlow™ in the assessment of fetal growth restriction – a randomized trial and prospective cohort study
- Advantages of fully automated AI-enhanced algorithm (5D CNS+™) for generating a fetal neurosonogram in clinical routine
- Reference ranges for fetal tricuspid and mitral annular plane systolic excursions
- FetalDenseNet: multi-scale deep learning for enhanced early detection of fetal anatomical planes in prenatal ultrasound
- Original Articles – Neonates
- Incidence and mortality trends in congenital diaphragmatic hernia in the United States
Articles in the same Issue
- Frontmatter
- Reviews
- Fetal neurobehavior and consciousness: a systematic review of 4D ultrasound evidence and ethical challenges
- Prenatal maternal stress and long-term neurodevelopmental outcomes: a narrative review
- From exencephaly to anencephaly: a catastrophic continuum of neural tube defects from embryogenesis to ultrasonographic diagnosis
- Opinion Papers
- Fetoception: a window into maternal interoception?
- Rationale for the use of fetal ventriculosubgaleal shunts for the treatment of aqueduct stenosis
- Original Articles – Obstetrics
- The fetal occiput-spine angle measurement during first stage of labor as a predictor for vaginal delivery, a systematic review and meta-analysis
- Hemorrhage-related maternal morbidity of secondary compared to primary postpartum hemorrhage
- First-trimester maternal serum PAPP-A levels and hyperemesis gravidarum: unraveling the link – a meta-analysis
- Antepartum cerebroplacental ratio in low risk pregnancy and its relationship with adverse perinatal outcome: a prospective cohort study
- Liver fibrosis markers as predictors of adverse outcomes in pregnancy-related hypertensive disorders
- Endocrine disrupting chemicals: translating mechanisms into perinatal risk assessment
- Low uterine segment thickness in the prediction of cesarean delivery after induction of labor
- Original Articles – Fetus
- Fetal adrenal gland size in preeclamptic pregnancies with and without fetal growth restriction: an ultrasonographic assessment
- Evaluation of safety and performance of CentaFlow™ in the assessment of fetal growth restriction – a randomized trial and prospective cohort study
- Advantages of fully automated AI-enhanced algorithm (5D CNS+™) for generating a fetal neurosonogram in clinical routine
- Reference ranges for fetal tricuspid and mitral annular plane systolic excursions
- FetalDenseNet: multi-scale deep learning for enhanced early detection of fetal anatomical planes in prenatal ultrasound
- Original Articles – Neonates
- Incidence and mortality trends in congenital diaphragmatic hernia in the United States

