Abstract
Proteasomes are the principal molecular machines for the regulated degradation of intracellular proteins. These self-compartmentalized macromolecular assemblies selectively degrade misfolded, mistranslated, damaged or otherwise unwanted proteins, and play a pivotal role in the maintenance of cellular proteostasis, in stress response, and numerous other processes of vital importance. Whereas the molecular architecture of the proteasome core particle (CP) is universally conserved, the unfoldase modules vary in overall structure, subunit complexity, and regulatory principles. Proteasomal unfoldases are AAA+ ATPases (ATPases associated with a variety of cellular activities) that unfold protein substrates, and translocate them into the CP for degradation. In this review, we summarize the current state of knowledge about proteasome – unfoldase systems in bacteria, archaea, and eukaryotes, the three domains of life.
Introduction
Selective protein degradation plays a crucial role in almost every aspect of cellular physiology. The degradation of mistranslated, misfolded, damaged or otherwise malfunctioning proteins is an essential element of protein quality control (Chen et al., 2011). Likewise, the timely removal of regulatory proteins is of vital importance in housekeeping functions such as cell cycle control, signal transduction, transcription, DNA repair and translation (Alves dos Santos et al., 2001; Goldberg, 2007; Bader and Steller, 2009; Koepp, 2014). Consequently, any disruption of selective protein degradation pathways leads to a broad array of pathological states, including cancer, neurodegeneration, immune-related disorders, cardiomyopathies, liver and gastrointestinal disorders, and ageing (Dahlmann, 2007; Motegi et al., 2009; Dantuma and Bott, 2014; Schmidt and Finley, 2014).
In eukaryotes, two major pathways have been identified for the selective removal of unwanted proteins – the ubiquitin-proteasome-system (UPS), and the autophagy-lysosome pathway (Ciechanover, 2005; Dikic, 2017). UPS constitutes the principal degradation route for intracellular proteins, whereas cellular organelles, cell-surface proteins, and invading pathogens are mostly degraded via autophagy. The two pathways significantly differ with respect to their substrates, and the machinery involved. However, the elements of control are carefully orchestrated, and jointly they ensure cellular protein homeostasis (Korolchuk et al., 2010).
The unifying element that links both UPS and selective autophagy is a small protein, ubiquitin, which marks substrates for destruction. Via an enzymatic cascade, ubiquitin is covalently bound to substrate proteins by an isopeptide bond between the carboxyl group of its C-terminal glycine residue, and the ε-NH2 group of substrate lysines. Iterations of the same reaction generate a ‘ubiquitin code’ of signals with different topologies and lengths (Ravid and Hochstrasser, 2008; Komander and Rape, 2012). Depending on the type of ubiquitin modification, a protein substrate is degraded either by the UPS or by autophagy (Kim et al., 2008; Korolchuk et al., 2010). Hence, ubiquitin labeling imparts specificity to a degradation process that is executed by an essentially non-specific molecular machine.
In bacteria and archaea, the existence of UPS or autophagy-lysosome pathways have not been reported. However, the occurrence of proteasomes, and ubiquitin-like small modifier proteins (SAMP and Pup) has been confirmed for all three domains of life.
Occurrence of proteasomes
The first description of proteasomes dates back to the late nineteen sixties, when a ‘cylinder shaped’ particle of unknown function was observed on electron micrographs of erythrocyte ghosts (Harris, 1968). More than a decade later, a cation-sensitive high-molecular-mass endopeptidase was discovered in bovine pituitaries, and designated the ‘multicatalytic protease complex’ (Wilk and Orlowski, 1980). Thereafter, the particle was rediscovered several times, and shown to exist ubiquitously in all eukaryotic cells. However, a lack of consensus over its biochemical nature, and physiological role, resulted in a plethora of names for the same protein complex. Eventually, the name ‘proteasome’ was coined (Arrigo et al., 1988), to highlight its character as a complex molecular machine with proteolytic function. For a review of the early history of the field, see Coux et al., 1996; Baumeister et al., 1997.
The first non-eukaryotic proteasomes were discovered in Thermoplasma acidophilum (Dahlmann et al., 1989), paving the way for a detailed analysis of proteasome structure and function in the other domains of life. Eventually, homologous protein complexes were isolated from Pyrococcus furiosus (Bauer et al., 1997), Methanosarcina thermophila (Maupin-Furlow et al., 1998), and several other archaea. Similarly in bacteria, the discovery of proteasomes in Rhodococcus erythropolis (Tamura et al., 1995) lead to its purification from Mycobacterium smegmatis (Knipfer and Shrader, 1997) and other related actinobacteria. So far, proteasomes have been characterized from at least 75 different genera, spanning all three domains of life (Figure 1).
![Figure 1: Dendrogram showing the distribution and relatedness of CP across domains of life.
The UPGMA tree is based on the cleaned and concatenated alignment of proteasome α and β sequences from species with fully sequenced genomes. In the case of genera with multiple α and β subunits, or with multiple fully sequenced species, a single representative member is chosen. To emphasize the overall similarity, structures of CP from Saccharomyces cerevisiae [PDB: 6FVW (Eisele et al., 2018)], Homo sapiens [PDB: 5L4G (Schweitzer et al., 2016)], T. acidophilum [PDB: 5VY3 (Herzik et al., 2017)], A. fulgidus [PDB: 6HE7 (Majumder et al., 2019)], Mycobacterium tuberculosis [PDB: 6BGL (Hu et al., 2018)] and R. erythropolis [PDB: 1Q5Q (Kwon et al., 2004)] are depicted surrounding the dendrogram. In each of the structures, identical subunits are colored identically.](/document/doi/10.1515/hsz-2019-0344/asset/graphic/j_hsz-2019-0344_fig_001.jpg)
Dendrogram showing the distribution and relatedness of CP across domains of life.
The UPGMA tree is based on the cleaned and concatenated alignment of proteasome α and β sequences from species with fully sequenced genomes. In the case of genera with multiple α and β subunits, or with multiple fully sequenced species, a single representative member is chosen. To emphasize the overall similarity, structures of CP from Saccharomyces cerevisiae [PDB: 6FVW (Eisele et al., 2018)], Homo sapiens [PDB: 5L4G (Schweitzer et al., 2016)], T. acidophilum [PDB: 5VY3 (Herzik et al., 2017)], A. fulgidus [PDB: 6HE7 (Majumder et al., 2019)], Mycobacterium tuberculosis [PDB: 6BGL (Hu et al., 2018)] and R. erythropolis [PDB: 1Q5Q (Kwon et al., 2004)] are depicted surrounding the dendrogram. In each of the structures, identical subunits are colored identically.
In eukaryotes, proteasomes are the only known soluble ATP-dependent proteases present in the cytoplasm and nucleus. In bacteria however, a more diverse set of ATP-dependent proteases (ClpXP, ClpAP, ClpCP, HslUV, Lon and FtsH systems) are known to exist, whose molecular architecture are analogous to the proteasome (Gur et al., 2011). Among these proteases, Lon is widespread, while an analogue of ClpP shows an infrequent presence in archaea (Maupin-Furlow, 2018). In eukaryotes, homologues of Lon, ClpP, and FtsH are only found in the cellular organelles of bacterial descent, such as mitochondria, and chloroplasts (Adam et al., 2001).
Structure of the proteasome core is universally conserved
Across the domains of life, proteasomes vary in subunit composition. However, the central component of all proteasomes, the core particle (CP) (also known as the 20S proteasome) exhibits a high level of architectural conservation (Figure 1). The CP is a barrel-shaped, self-compartmentalized, protein complex, composed of four stacked seven-membered rings (Baumeister et al., 1998). The outer rings comprise α-type subunits, whereas β-type subunits form the inner rings. The co-axial stacking of four rings (α7β7β7α7) creates three internal cavities, bound by four narrow constrictions, as originally observed in three-dimensional (3D) reconstructions from electron micrographs (Hegerl et al., 1991). The central cavity, formed at the junction of two β-rings, is the catalytic chamber, where protein degradation takes place. The two outer cavities, formed at α-β junctions serve as antechambers, where substrate proteins can be stored in an unfolded state prior to degradation (Sharon et al., 2006; Ruschak et al., 2010).
In archaea and actinobacteria, only one or two types of α and β-subunits exist, while in eukaryotes, at least 14 unique subunits (α1-7 and β1-7) build the constitutive CP. In addition, higher eukaryotes express specialized proteasomes such as the immunoproteasomes and thymoproteasomes, where β subunit variants (β1i, β2i, β5i, and β5t) substitute the house-keeping ones (Murata et al., 2018). The α- and β-type subunits are structurally related, differing only in their N-terminal regions. The N-terminal tails of α-subunits form a CP gate, which prevents access of folded proteins into the CP-axial channel (Wenzel and Baumeister 1995). Whereas the N-terminal regions of β subunits serve as pro-peptides that are autocatalytically cleaved off during proteasome maturation to expose the proteolytic active sites (Seemuller et al., 1996; Groll et al., 1999; Huber et al., 2016).
Proteasome active sites are sequestered within the CP
Active sites of the proteasome are located on the inner walls of the CP catalytic chamber, and comprise a triad of residues Thr1, Lys33, and Asp/Glu17 (Huber et al., 2016) (Figure 2). These residues are conserved among all proteolytically active β-type subunits in eukaryotic, bacterial and archaeal proteasomes, and hence the mechanism of peptide bond cleavage follows a universal principle among all CPs (Figure 2). The substrate cleavage preference is however distinct for every subunit, and depends on the chemical nature of the substrate-binding channel, especially the active site’s S1 specificity pocket (Groll and Huber, 2003). Thus, the CP in archaea and actinobacteria comprise 14 identical catalytic sites (Seemuller et al., 1995; Tamura et al., 1995). Whereas in eukaryotes, proteolytic activity is restricted to only β1, β2, and β5 type subunits (including β1i, β2i, β5i, β5t), and hence the CP harbors a maximum of six catalytic sites, with three distinct peptide cleavage specificities (Orlowski, 1990; Seemuller et al., 1995; Heinemeyer et al., 1997) (Figure 2).

CP β-subunits harbor the proteolytic active sites.
(A) Dendrogram showing the relatedness of an aligned set of β-type subunit sequences from Ta: T. acidophilum, Af: A. fulgidus, Mj: Methanocaldococcus jannaschii, Mt: M. tuberculosis, Re: R. erythropolis, Sc: S. cereviseae, Bt: Bos taurus, Hs: Homo sapiens, Mm: Mus musculus, and Rn: Rattus norvegicus. The selection of organisms is based on the availability of structural information (PDB entries), and the protein names are consistent with their respective Uniprot Gene IDs. Inactive β-type subunits are shown in gray, while active β-type subunits from constitutive proteasomes are shown in green. Specialized β-type subunits of thymoproteasomes and immunoproteasomes are indicated in yellow and blue, respectively. (B) Close-up views of the proteolytic active sites of a human constitutive CP and a T. acidophilum CP. The active site surfaces in each case are colored according to their charge distribution in a scale indicated by the key below. Conserved residues comprising the catalytic triad (T1, D/E17, and K33) are shown in black, while other residues identified that contribute to the charge landscape of the substrate-binding channel, are indicated in green. Contrary to the prevailing views at the time, the residues constituting the ‘catalytic triad’ turned out to be the same for the ‘trypsin-like’, ‘chymotrypsin-like’ and ‘PGPH-like’ activities, and specificity happens to be conferred by the local environment of the substrate-binding channel.
Proteasomes are classified as a family of N-terminal nucleophilic (Ntn) threonine proteases, where the free N-terminal Thr1 of a catalytically active β-subunit acts as a nucleophile in peptide bond hydrolysis (Seemuller et al., 1995). For mechanistic details, see Huber et al., 2016. Back in the day, the discovery of this unique mechanism immediately suggested a way of inhibiting the CP by capping the β-subunit N-termini, and in turn inspired the development of an important therapeutic agent bortezomib. For a brief history of the development of proteasome inhibitors, see Goldberg, 2012.
Like many proteases, the proteasome β-subunits are synthesized in an inactive precursor form. Pro-peptides of varying lengths cap the active site prior to proteasome maturation and activation. These pro-peptides are believed to help drive the proper assembly of the CP, and simultaneously prevent the irreversible inactivation of the Thr1 N terminus by N-acetylation (Arendt and Hochstrasser, 1999; Groll et al., 1999; Jager et al., 1999; Kwon et al., 2004). For a detailed account of the CP assembly process, and the chaperones involved, see Zuhl et al., 1997; Murata et al., 2009; Kunjappu and Hochstrasser, 2014.
Substrate processing requires proteasome activation
Protein degradation by the CP does not require ATP hydrolysis as such. However, the closed compartmentalized structure of the CP limits its constitutive proteolytic activity to only denatured or unstructured proteins and small peptides (Wenzel and Baumeister, 1995; Liu et al., 2003). Larger protein substrates are mostly degraded via ‘unfoldase-assisted-proteolysis’, whereby the CP recruits cofactors or activators to facilitate substrate access into its catalytic chamber.
Several proteasomal activators have thus far been identified, spanning both ATP-dependent and independent varieties (Stadtmueller and Hill, 2011). These activators cap the CP at one or both ends, and reposition the α-ring gating residues into an open conformation. For the processing of folded proteins however, gate opening is not sufficient to allow access into the catalytic chamber. Additional steps of substrate unfolding and translocation are required, and can be achieved only at the cost of ATP hydrolysis. As a result, proteolysis is rendered an energy-dependent process.
Energy-dependent proteasome activators
In all domains of life, energy-dependent proteasome activation involves ATPases of the AAA+ variety (Yedidi et al., 2017). These are an extended group of ATPases associated with various cellular activities, among which AAA-ATPases form a sub-group. Members of the AAA+ superfamily are defined by the presence of AAA+ domains that use six conserved motifs for function; Walker A, Walker B, sensor-1, arginine finger (R-finger), sensor-2 and pore loops (Wendler et al., 2012; Miller and Enemark, 2016). The Walker A and Walker B motifs are required for ATP binding and hydrolysis. The sensor-1 and R-fingers coordinate nucleotide hydrolysis and propagate the associated conformational changes between subunits (Ogura et al., 2004). The sensor-2 region mediates conformational changes associated with a cycle of ATP binding and hydrolysis, whereas the pore loops make direct contact with substrate (Martin et al., 2008).
Proteasomal ATPases share a common architecture and assemble into hexameric ring complexes, with N-terminal domains, AAA+ domains, and C-terminal HbYX motifs (where Hb is a hydrophobic residue, Y is tyrosine, and X is any amino acid). The N-terminal domain is believed to be involved in substrate capture. The HbYX motif is responsible for interaction with the CP, while the AAA+ domain unfolds protein substrates, and translocates them through the axial channel, into the degradation chamber. Such an association of hexameric ATPases with a heptameric CP α-ring gives rise to ‘symmetry-mismatched’ complexes.
In eukaryotes, the proteasomal ATPase is an assemblage of ‘regulatory particle triple A ATPases’ (Rpt1-6) that forms the core of the 19S regulatory particle (RP). In archaea however, the proteasome is believed to operate via a network of AAA+ ATPases (Forouzan et al., 2012), among which the ‘proteasome activating nucleotidase’ (PAN), and the Cdc48 homologue ‘VCP-like ATPase’ (VAT) have been characterized. For a long time only one proteasomal ATPase was identified in actinobacteria, the ‘AAA-ATPase forming ring-shaped complex’ (ARC) (Wolf et al., 1998), which is known as the ‘mycobacterial proteasome ATPase’ (Mpa) in Mycobacterium (Darwin et al., 2005). Recently however, a Cdc48 homologue ‘Cdc48-like protein of actinobacteria’ (Cpa) has also been observed to interact with the proteasome in a manner similar to other proteasomal ATPases (Ziemski et al., 2018).
Similar to the CP, the subunit complexity of proteasomal ATPases varies in the different domains of life. Thus, the archaeal or actinobacterial ATPases are homohexameric, whereas the Rpts of the 26S proteasome form a heterohexameric complex, which is further associated with non-ATPase subunits (Rpns) in the RP. Due to the absence of Rpn-equivalents, non-eukaryotic proteasomes presumably function as minimal complexes, where the ATPases alone are responsible for substrate capture, in addition to CP activation and substrate unfolding. However, these CP-ATPase complexes are rather unstable, and notoriously difficult to isolate from cell lysates. Hence, a question is often raised regarding the constitution and stability of these proteasomes, hinting at a fleeting interaction between CP and ATPase, or the involvement of additional stabilizing co-factors that are yet unknown.
PAN
PAN was the first proteasome-activating AAA-ATPase discovered in archaea (Zwickl et al., 1999). It exhibits ~41–45% sequence homology to the Rpts of the 26S proteasome (Zwickl et al., 1999), and is considered as its evolutionary precursor. In terms of domain architecture, it is the simplest among proteasomal ATPases, and comprises an N-terminal helix, an oligonucleotide/oligosaccharide binding (OB) domain, an ATPase domain, and a C-terminal HbYX motif. The association of six PAN protomers creates three two-stranded coiled-coils, an OB-ring and an AAA-ring, all of which are connected via flexible linkers (Djuranovic et al., 2009; Zhang et al., 2009a).
Biochemical studies over the past two decades have confirmed a chaperone activity of PAN, by which it is able to prevent the in vitro aggregation of denatured proteins, and assist in their refolding (Benaroudj and Goldberg, 2000; Navon and Goldberg, 2001). There have also been speculations regarding the conformational switching of N-domains (Djuranovic et al., 2009; Snoberger et al., 2018), nucleotide usage by the AAA-ring (Zhang et al., 2009b; Smith et al., 2011; Kim et al., 2015), and the involvement of HbYX tails in CP gate opening (Smith et al., 2007; Rabl et al., 2008). However, the physical association between PAN and CP has been shown for the first time in 3D in our recent cryo-EM structure of the Archaeoglobus fulgidus PAN-proteasome (Majumder et al., 2019).
Under in vitro conditions, the association of PAN and CP requires ATP. The interaction is short-lived, and PAN-proteasome holocomplexes are enriched only in the presence of the slowly hydrolyzable nucleotide analog ATPγS (Smith et al., 2005). Within a PAN-proteasome, PAN interacts with the CP via its C-terminus, while the N-terminal coiled-coils are free (Figure 3). A clear offset is visible between PAN and CP, and their axial channels are not perfectly aligned. However, the PAN AAA-ring adopts a uniform spiral staircase conformation, with helically arranged pore loops protruding into the axial channel.

Cryo-EM structure of the A. fulgidus PAN-proteasome (Majumder et al., 2019).
(A) Architecture of a representative (state 2) PAN-proteasome pseudo-single-capped particle (PSC). Model of the PAN-proteasome in state 2 (PDB: 6HE9) is superimposed on the respective PSC density (EMDB: EMD-0213). Each PAN protomer is colored differently, while the CP is in gray. (B) PAN binding stabilizes the CP gate in an open conformation. In the absence of PAN, the free CP possesses an unstructured gate, which appears as a smeared density occupying the axial pore. The inter α-subunit pockets remain un-occupied. Whereas, in the case of the PAN-bound CP, the axial channel is clear, with finger-like projections surrounding the axial pore. The finger-like projections correspond to the N-terminal tails of CP α-subunits that have stiffened upon PAN-binding. HbYX insertions are visible within inter α-subunit pockets. Five HbYX insertions show strong density, while the sixth one is weak. (C) Structure-based model for the PAN-proteasome ATPase cycle. Densities and superimposed models are shown for the five conformational states identified, alongside their respective cartoon representations. The motion of the OB-ring is emphasized by black lines. Within the scheme, the subunits in the highest and lowest position of the AAA-staircase are indicated with colored ‘H’ and ‘L’, respectively, and the active (ADP) pocket is highlighted with asterisks.
Interestingly, the interaction between PAN and CP is bipartite. While the insertion of C-terminal HbYX tails into inter α-subunit pockets establishes a classical contact between PAN and CP, the N-terminal residues of the CP α-subunits extend outwards to engage in ‘gate contacts’ with the lowest protomer in the PAN AAA-staircase.
Extensive in silico classification allowed us to deconvolute five conformational states coexisting in the sample. The conformational states show the sequential rotation of the PAN AAA-staircase (Figure 3). In each state, PAN is positioned at an offset atop the CP axial channel, yet the CP exhibits an open gate with all six PAN HbYX tails inserted within inter α-subunit pockets. The PAN AAA-staircase features co-existing nucleotide states, and correlated nucleotide-dependent inter-subunit signaling contacts between adjacent protomers. The OB-ring and coiled-coils that seem to perch on top of the AAA-ring, also adjust with the conformational changes in the AAA-ring, and slant from the highest protomer.
Guided by the structural information from the five conformational states, it appears that progress of the ATP hydrolysis cycle is accompanied by the making and breaking of inter-subunit contacts, which in turn causes the rotation of the AAA-staircase. Thus, a sequential ATP hydrolysis event leads to a harmonic up and down motion of the pore helices and pore loops around the AAA-ring. This motion is communicated via the linkers to the OB ring and coiled coils, and helps drive the gripping, unfolding, and translocation of substrate proteins into the CP.
VAT
VAT is an archaeal homologue of the widely occurring Cdc48/p97/VCP ATPase (Golbik et al., 1999; Gerega et al., 2005), which is ubiquitous in both archaea and eukaryotes. Cdc48 plays a crucial role in a wide range of cellular activities (Meyer et al., 2012), including the unfolding of ubiquitin (Twomey et al., 2019). VAT was recognized as a proteasome activator when PAN was found not to exist in some archaea, and not required for the viability of some others, where the CP was essential (Zhou et al., 2008). Interestingly, the involvement of Cdc48-type proteins in proteasome activation has thus far been reported only for archaea and actinobacteria (Barthelme and Sauer, 2012; Forouzan et al., 2012; Ziemski et al., 2018). In eukaryotes, a collaboration between Cdc48/p97 and 26S proteasomes has been implicated, particularly in the context of endoplasmic-reticulum-associated protein degradation (ERAD) (Bodnar and Rapoport, 2017). However, compelling evidence for a direct physical interaction between Cdc48 and proteasomes is still lacking. The prevailing view is that p97, together with Ufd1/Npl4, extracts polyubiquitinated proteins from the ER, and hands them over to the proteasome (Wu and Rapoport, 2018). However, in situ studies have not shown a direct or stable interaction between Cdc48 and 26S proteasomes.
VAT is a Janus-faced chaperone occurring in the archaeon T. acidophilum (Pamnani et al., 1997; Golbik et al., 1999). Depending on the concentration of Mg2+ present, VAT assumes either a low activity state that is capable of refolding proteins or a high activity one, which accelerates the unfolding of the same (Golbik et al., 1999).
It is a member of the Type II family of AAA+ ATPases, in which each protomer comprises three domains: an N-terminal domain that is divergent between AAA+ proteins, and two tandem nucleotide binding domains. In presence of the slowly hydrolyzable nucleotide analogue ATPγS, full length VAT exists either as a six-fold symmetric stack of rings, or as helical split-rings, the latter conformation being predominant also in presence of ADP (Huang et al., 2016) (Figure 4). Typical of proteasomal ATPases, the VAT C-terminus contains an HbYX motif, which is involved in an interaction with the CP. The nucleotide binding domains coordinate ATP hydrolysis and substrate unfolding. However, the role of the N-terminal domain is unclear, and as suggested by mutational studies, it might partially inhibit the unfolding activity of VAT (Gerega et al., 2005). In fact, the N-terminal deletion mutants of VAT are more stable, and show increased CP binding affinity compared to full-length ones (Barthelme and Sauer 2012).

Substrate processing by the T. acidophilum VATΔN (Ripstein et al., 2017).
(A) Cryo-EM densities and superimposed models of the ATPγS bound VATΔN (PDB: 5VC7) is shown from two different angles. It has a characteristic six-fold symmetry, with a central pore/axial channel running through the middle of the complex. (B) The structure of the substrate-bound VAT complex (PDB: 5VCA). The substrate density is visible only in filtered versions of the density map (EMDB: EMD-8659), and its trajectory through the VAT complex is emphasized by the red dashed line. In each case, the N-terminal nucleotide binding domain (NBD1) is shown in sand, and the C-terminal nucleotide binding domain (NBD2) is colored dark green. Upon substrate binding, a conformational change in the NBD rings leads to a helical architecture.
Pull-down assays indicate a short-lived interaction between VAT and CP (Barthelme and Sauer, 2012). In fact, the visualization of VAT-CP complexes has been possible only after site-specific crosslinking (Barthelme et al., 2014), although VAT has a higher affinity for the CP [Kd(VAT-CP)=160 nm] compared to PAN [Kd(PAN-CP)=~300 nm] (Barthelme and Sauer, 2012). Nevertheless, cryo-EM and NMR studies of substrate bound VAT have provided invaluable insights into its mechanism of unfolding and translocation (Ripstein et al., 2017; Augustyniak and Kay, 2018).
Trapping calmodulin bound VAT via engineered inter-molecular disulfide bridges, it was shown that VAT initiates substrate translocation by pulling it onto intrinsically unstructured regions, irrespective of amino acid sequence, or location (Augustyniak and Kay, 2018). Substrate binding is accompanied by a conformational change in each protomer of N-terminal deleted VAT, so as to rearrange the nucleotide binding domains from a stacked-ring conformation into a split lockwasher (Figure 4). The transition occurs via the horizontal displacement and rotation of one nucleotide binding domain with respect to the other, and results in a decreased distance between their pore loops. The pore loops of five of six protomers interact with the substrate, wrapping ~80 Å length of substrate in a hydrophobic helical sheath. The sixth (non-interacting) protomer, occupies the top and bottom of the helix, forming a seam subunit with dramatically distinct protomer-protomer interactions. Thus, by a processive ‘hand-over-hand’ mechanism, substrates are translocated as each VAT protomer releases its grip on the substrate, and re-engages further along the target (Ripstein et al., 2017).
A new addition to the list of Cdc48-type proteasome activators is Cpa, which by phylogenetic analysis appears to descend from a common Cdc48 ancestor (Ziemski et al., 2018; Müller and Weber-Ban, 2019). The interaction of Cpa with the proteasome has been recently recognized, and remains to be explored in detail. Like the other proteasomal AAA+ ATPases, Cpa forms hexameric ring complexes. However, its oligomerization occurs only presence of nucleotide. Interestingly, Cpa lacks a classical HbYX motif, and yet binds the CP with an affinity [Kd(Cpa-CP)=360 nm] comparable to a PAN-CP interaction [Kd(PAN-CP)=~300 nm].
MPA
In actinobacteria, proteolysis by the CP requires assistance of the AAA-ATPase ARC/MPA (Darwin et al., 2005). The MPA-proteasome system renders Mycobacterium tuberculosis resistant to killing by host macrophages, and plays a crucial role in tuberculosis (Samanovic and Darwin, 2016). Consequently, the MPA-proteasome system serves as an attractive target for the development of anti-tuberculosis therapeutics (Lupoli et al., 2018).
ARC/MPA shares structural similarity with the archaeal PAN or the eukaryotic Rpts. Each MPA protomer comprises an N-terminal extended α-helical region, followed by two tandem oligonucleotide/oligosaccharide binding (OB) domains, an AAA-domain, a β-grasp domain and a C-terminal GQYL motif (Wu et al., 2017). Upon oligomerization, adjacent α-helical domains form three pairs of two stranded coiled-coils, while the OB and AAA domains assemble into co-axially stacked hexameric rings.
The N-terminal coiled-coils in MPA serve as a specific receptor for the small ‘prokaryotic ubiquitin-like protein’ (Pup) (Sutter et al., 2009). Pup, which is an otherwise unstructured protein, interacts with the coiled-coils in MPA via its N-terminus and assumes a helical structure (Wang et al., 2010). It is believed that MPA-proteasome complexes engage Pup-tagged substrates, and unfolds and drags them into the CP (Striebel et al., 2010). The precise role of the OB-domains is unclear. However, they have been found sufficient to maintain the hexameric state of MPA (Wang et al., 2009). The C-terminal GQYL motif is a functional equivalent of the HbYX motif found in other proteasomal ATPases, and is believed to be responsible for activator-CP interaction (Bolten et al., 2016; Hu et al., 2018). However, the β-grasp domain buries the C-terminus of each protomer in the central channel of the MPA hexamer, and hinders interaction with the CP (Wu et al., 2017).
Incidentally, a 3D structure of an MPA-proteasome complex is still lacking. Only a loose physical association between MPA and CP has thus far been observed, and is limited to gate-deleted CP constructs (Wang et al., 2009). In fact, MPA alone cannot facilitate robust in vitro degradation by the CP, and requires the presence of additional factors such as the proteasome accessory factor (PafA or PafE), a Pup ligase, a depupylase (Dop), and possibly others that are yet unknown (Pearce et al., 2008; Striebel et al., 2009; Burns et al., 2010; Hu et al., 2018).
19S RP
Among the proteasome activators known thus far, the 19S RP is the most complex macromolecular assemblage. It forms the regulatory component of the 26S proteasome, and is composed of at least 19 canonical subunits, along with an array of substoichiometric ‘proteasome interacting proteins’ (Pips). The RP can be biochemically separated into the ‘base’ and ‘lid’ subcomplexes (Glickman et al., 1998). Such a separation assigns 10 subunits to the base, among which are six distinct AAA+ ATPases (Rpt1-6), and four non-ATPase subunits (Rpn1, Rpn2, Rpn10 and Rpn13). The remaining nine subunits (Rpn3, Rpn5, Rpn6, Rpn7, Rpn9, Rpn12, Rpn8, Rpn11, and Sem1) constitute the lid. For a detailed structural description of the 19S RP, see Beck et al., 2012; Lander et al., 2012; Lasker et al., 2012 and Schweitzer et al., 2016 (Figure 5).
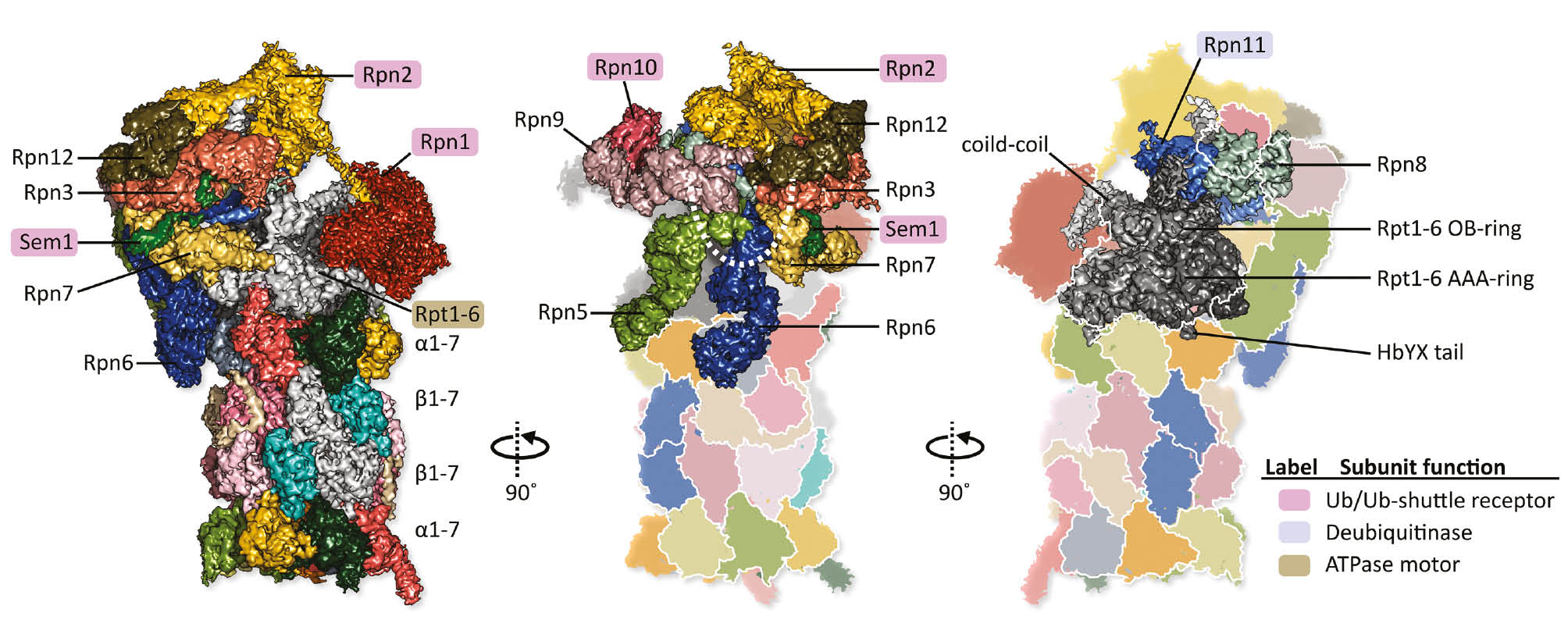
Cryo-EM structure of a single-capped human 26S proteasome is shown in three rotated orientations (EMDB: EMD-4002; Schweitzer et al., 2016).
The CP is formed of four stacked hetero-heptameric rings (α1-7 β1-7 β1-7 α1-7), whereas the RP comprises a hetero-hexameric ATPase (Rpt1-6) core, surrounded by non-ATPase subunits (Rpn1-3, Rpn5-13, and Sem1). Highlighted labels indicate the RP subunits with specialized roles in substrate processing. The corresponding functions are enlisted in the color key on the right. Ubiquitin receptor subunits are peripherally located on the RP. Rpn10 and Rpn13 are the primary ubiquitin receptors, while Rpn1, Rpn2 and Sem1 recruit ubiquitinated substrates via shuttle factors. In the human 26S proteasome preparations, Rpn13 is present in substoichiometric amounts, and therefore missing in the density map. However, it is known to be recruited via Rpn2. The view in the middle exhibits the proteasome lid with a horseshoe-like arrangement of scaffolding subunits (Rpn12-Rpn3-Rpn7-Rpn6-Rpn5-Rpn9). These subunits presumably play structural roles, with Rpn6 clamping together the RP and the CP. The view on the right highlights the Rpt1-6 motor (in shades of gray). It comprises an AAA-ring, an OB-ring, and three coiled-coils. The Rpt1-6 AAA-ring docks onto the CP via insertion of C-terminal tails (of Rpt1, Rpt2, Rpt6, Rpt3, and Rpt5) into inter α-subunit pockets. The insertion of Rpt1 and Rpt6 C-termini lead to CP gate opening. Substrates are deubiquitylated by Rpn11, which forms a hetero-dimer with Rpn8, and is located close to the Rpt1-6 OB-ring. The other proteasomal DUBs, Ubp6 (recruited via Rpn1) and Uch37 (recruited via Rpn13), could not be localized in the density map.
At the core of the 19S RP is a hexameric assemblage of Rpt1-2-6-3-4-5 ATPases (Tomko et al., 2010), which serves as the molecular motor of the 26S proteasome. It resembles the archaeal PAN in overall structure, and comprises an AAA-ring, an oligosaccharide/oligonucleotide binding (OB) ring (also known as the N-ring), and coiled-coils between pairs of neighboring subunits (Rpt1/Rpt2, Rpt6/Rpt3, and Rpt4/Rpt5). The Rpt1-6 motor is primarily responsible for substrate unfolding and translocation into the CP. In addition, it regulates CP gate opening by insertion of its C-terminal HbYX tails into the CP inter α-subunit pockets. The remaining subunits, which include ubiquitin receptors, deubiquitylases (DUBs), and scaffolding proteins, partially embrace the Rpt1-6 hexamer and the CP. The resulting physical interaction between ATPase and non-ATPase subunits helps to connect substrate recruitment and ubiquitin processing to the unfolding machinery.
Ubiquitin receptors are placed along the periphery of the RP, where they are accessible to protein substrates. Being highly flexible, they are the least well-resolved parts in 26S proteasome structures. Rpn10 binds polyubiquitin chains through its ubiquitin-interacting motif (UIM), while its N-terminal von Willebrand A (VWA) motif mediates its interaction with the proteasome (Deveraux et al., 1994; Verma et al., 2004; Wang et al., 2005). The yeast Rpn10 contains a single UIM. However, the number of UIMs increase in higher eukaryotes, and the additional UIMs can serve as binding sites for ubiquitin-like (UBL) domain containing shuttle factors. It has been shown that Rpn10 functions both when free, and in the proteasome bound form (Matiuhin et al., 2008). However, ubiquitylation of Rpn10 regulates its capacity to further bind ubiquitinated substrates (Isasa et al., 2010).
Rpn13 recognizes ubiquitinated substrates through its N-terminal pleckstrin-like receptor of ubiquitin (PRU) domain, while its opposite surface interacts with Rpn2 (Husnjak et al., 2008; Lu et al., 2017). In higher eukaryotes, Rpn13 contains an additional DEUBiquitinase adaptor (DEUBAD) domain, which interacts with a DUB Uch37 (Hamazaki et al., 2006; Yao et al., 2006). Like Rpn10, Rpn13 also binds ubiquitin-like (UBL) domains of UBL-ubiquitin-associated (UBA) proteins. The combination of ubiquitin binding and deubiquitinating functionalities renders Rpn13 as a ‘proof-reading’ machinery that enables substrates to either engage or escape from the proteasome.
Rpn1, a PC-repeat protein, serves as a ligand-binding hotspot, with two adjacent receptor sites for substrate binding and deubiquitination (Shi et al., 2016). With the capacity to recognize both ubiquitin, and UBL domains of substrate shuttling factors, Rpn1 recruits extrinsic ubiquitin shuttle receptors (Rad23, Ddi1 and Dsk2) and the non-essential DUB Ubp6 to the proteasome (Elsasser et al., 2002; Leggett et al., 2002; Gomez et al., 2011). Ubiquitin recognizing properties have also been associated with the scaffolding protein Sem1/Dss1 (Paraskevopoulos et al., 2014), and the proteasomal ATPase, Rpt5 (Lam et al., 2002), however these are less well characterized.
The 19S RP contains three DUBs – Rpn11, Ubp6/Usp14 and Uch37, among which only Rpn11 is essential. It is a Zn2+ containing JAMM metalloprotease, which forms an obligate heterodimer with its neighboring subunit Rpn8. Rpn11 exists in a self-inhibited state (Pathare et al., 2014; Worden et al., 2014), which is further stabilized by its neighboring lid subunit, Rpn5 (Dambacher et al., 2016). In the ground state of the proteasome, it is located above the Rpt1-6 motor, between Rpn2 and the ubiquitin receptor Rpn10. Upon substrate engagement, a conformational switching brings Rpn11 to the mouth of the substrate translocation channel (Matyskiela et al., 2013), and substrate-bound ubiquitin moieties are directly pulled into its catalytic groove (de la Pena et al., 2018).
The two other DUBs, Ubp6/Usp14 and Uch37, associate with the RP in substoichiometric amounts. Ubp6 tethers to Rpn1 via an N-terminal UBL domain, whereas its C-terminal catalytic domain is positioned variably (Aufderheide et al., 2015). Interestingly, ubiquitin docking to the active site of Ubp6 ‘non-catalytically’ inhibits protein degradation by the proteasome (Bashore et al., 2015; Kim and Goldberg 2017). The other DUB Uch37, associates with proteasomes via interaction of its C-terminal Uch37-like domain (ULD) and the DEUBAD of Rpn13 (Yao et al., 2006; Vander Linden et al., 2015). Like Rpn11, Uch37 exists in a self-inhibited state, and binding to RPN13 relieves the inhibitory effect (Yao et al., 2006; Jiao et al., 2014).
The remaining 19S RP subunits are believed to play structural roles. Rpn3, Rpn7, Rpn6, Rpn5, Rpn9 and Rpn12 are arranged in a horseshoe-shaped anchor from which their N-terminal domains extend radially. These subunits are characterized by a C-terminal PCI (proteasome – cop9 signalosome – initiation factor 3) domain, and likely act as scaffolds (Finley 2009). Rpn2, which is structurally similar to Rpn1, also functions as a scaffold, binding Rpn13 and ubiquitin shuttles (Rosenzweig et al., 2012). Due to the presence of these structural subunits, especially the RP-CP clamping subunit Rpn6 (Pathare et al., 2012), the 26S proteasome is significantly more stable than the proteasomal ‘minimal complexes’ found in the other domains of life. In fact, intact endogenous 26S proteasomes are routinely purified from eukaryotic cell lysates, and have also been successfully visualized in situ (Asano et al., 2015; Albert et al., 2017; Guo et al., 2018).
Mechanism of unfoldase- assisted-proteolysis
Thanks to the numerous cryo-EM studies on yeast and human 26S proteasomes, the proteasome conformational landscape has been significantly expanded in recent years (Matyskiela et al., 2013; Sledz et al., 2013; Ding et al., 2017; Wehmer et al., 2017; de la Pena et al., 2018; Eisele et al., 2018; Zhu et al., 2018; Ding et al., 2019; Dong et al., 2019). However, the increasing array of conformational states makes it challenging to predict the actual sequence of events. The conformations observed reflect large scale motions of the proteasome lid that lead to the relocation of certain RP subunits, and a concomitant restructuring of the inter-subunit interaction network. The latter is believed to play a vital role in ubiquitin recognition and processing, the properties unique to eukaryotic proteasomes. However, the unfolding and translocation of engaged substrates is controlled largely by the Rpt1-6.
Both yeast and human 26S proteasomes exhibit different planar and rotated spiral staircase conformations of the Rpt1-6 ring. This indicates the co-existence of multiple nucleotide states, or an ATPase cycle in progress. With the parallel discovery of rotated spiral staircases in the archaeal PAN, the fundamental role of an around-the-ring ATPase cycle is further established. Although the sequence of steps in an ATPase cycle is yet to be determined, the existence of inter-subunit signaling motifs in both PAN and Rpt1-6 indicate an underlying communication between adjacent protomers during the progression of the cycle. Another notable feature conserved between PAN and the Rpt1-6 is the motion of the OB ring, which is coordinated with the ATPase domain transitions. In the case of PAN, such motions lead to the wagging of the exposed N-terminal coiled-coils, like “swinging arms’, and are possibly responsible for substrate capture. However, in the case of Rpt1-6, motions of the OB-ring and coiled-coils results in varied interactions with lid subunits. As non-eukaryotic proteasomes require the assistance of AAA+ unfoldases, one would expect that such transitions in the Rpt1-6 motor in turn coordinate the conformational changes in the 26S proteasome lid.
The functional cycle of mycobacterial proteasomes however, seem to be distinct from the eukaryotic or archaeal counterparts. As the presence of a β-grasp domain in MPA hinders its HbYX motifs from physically interacting with the CP, the CP is presumably activated with the help of accessory factors PafA or PafE. It would be interesting to note how gate opening, substrate unfolding, and translocation are coordinated in such systems.
ATP-independent activators
In addition to the ATPase activators of the CP, several non-ATPase activators have been identified. They are less widely conserved, and their substrates and biological functions are less clear. However, in most cases, the interaction between CP and non-ATPase activators have been successfully characterized from a structural perspective (Forster et al., 2005; Sadre-Bazzaz et al., 2010).
In eukaryotes, ATP-independent proteasome activators belong to two major categories: PA200 and PA28 (11S). The PA200 activators in human, and their homologous Blm10 activators in yeast, are composed of a single polypeptide chain with an array of HEAT repeats. This elongated solenoid gives rise to a dome like architecture (Sadre-Bazzaz et al., 2010), which forms complexes with the CP (Schmidt et al., 2005; Toste Rego and da Fonseca, 2019). The C terminus of the Blm10 chain contains a YYX motif (analogous to the HbYX motif of ATP-dependent activators) that binds between the CP α5 and α6 subunits, and induces partial opening of the CP gate (Dange et al., 2011). These activators are believed to participate in a broad variety of processes, including CP assembly (Fehlker et al., 2003), DNA repair (Ustrell et al., 2002), genomic stability (Blickwedehl et al., 2008), proteasome inhibition (Lehmann et al., 2008), spermatogenesis (Khor et al., 2006), and mitochondrial function (Sadre-Bazzaz et al., 2010).
Higher eukaryotes and certain unicellular eukaryotes (such as trypanosomes) additionally express a PA28/11S/REG family of proteasome activators. However, they are missing in yeast and plants. The PA28 activators are heptameric toroids that form hybrid complexes with the CP (Hendil et al., 1998). Unlike the PA200/Blm10 activators, here each subunit of the heptamer interacts with the CP via internal activation loops, and non-HbYX type insertions of their C-terminal residues (Forster et al., 2005). In mammals, three different isoforms of PA28 have been identified (PA28α, β, and γ), among which PA28γ has earned an attention due to its role in cancer biology (He et al., 2012). PA28 activators are believed to be involved in cell cycle control and apoptosis (Murata et al., 1999; Masson et al., 2003), MHC class-I antigen presentation (Groettrup et al., 2010), and a variety of cellular roles.
ATP-independent proteasome activators have also been found in bacteria and in archaea. PafE (Bpa) is functionally similar but evolutionarily unrelated to the eukaryotic proteasome activators PA26 and PA28, and contributes to the virulence of M. tuberculosis (Jastrab et al., 2015). Cryo-EM studies have recently elucidated how a dodecameric assemblage of PafE interacts with the CP via its C-terminal GQYL motifs (Bolten et al., 2016; Hu et al., 2018). In archaea however, PbaA and PbaB are homologous to the eukaryotic CP assembly factors PAC1-PAC2. PbaB forms a homotetramer with tentacle-like C-terminal HbYX segments. It can function both as a molecular chaperone as well as an ATP-independent proteasome activator (Kumoi et al., 2013).
Concluding remarks
In the three domains of life, proteasomes and their activators show substantial structural and functional overlap, and yet there are key differences in their mechanisms of assembly, activation, and substrate targeting for degradation. Like the UPS in eukaryotes, the proteasomes in actinobacteria function as a part of the Pup-proteasome system, whereby substrates tagged with the small modifier protein Pup are recognized by MPA, and targeted to the proteasome (Pearce et al., 2008). Similarly, in archaea, the enzymatic attachment of a ubiquitin-like modifier protein SAMP (Humbard et al., 2010), serves as a signal for proteasome mediated proteolysis (Fu et al., 2016). Not to mention, both archaea and actinobacteria possess elaborate machineries for the conjugation and removal of Pup and SAMP, respectively.
In evolutionary terms, the bacterial ATP-dependent protease, HslUV, is considered a prototype of the proteasome. This two-component complex, composed of AAA+ (HslU), and proteolytic (HslV) units, is related to the proteasome both structurally and functionally (Rohrwild et al., 1996). With sequence homology between HslV and proteasome β subunits, HslUV even demonstrates a threonine-dependent proteolytic mechanism, typical of proteasomes. It is therefore tempting to investigate in greater depth how the proteasomal systems have evolved. However, efforts in this direction have not yet yielded a hypothesis, largely due to the limited knowledge of evolutionary precursors.
The recent identification of two proteasomal evolutionary intermediates, ancestral β subunit (Anbu), and betaproteobacterial homolog (BPH) have allowed the analysis of how transitions in symmetry mark pivot points in the evolution of the proteasome family (Fuchs et al., 2017, 2018; Fuchs and Hartmann, 2019). Characterization of the PAN-proteasome has also provided invaluable insights into the functional cycle of archaeal proteasomes (Majumder et al., 2019). Certain structural features such as ‘gate contacts’ have been revealed, which resemble the bipartite communication modes present in bacterial ClpXP (Martin et al., 2007). These contacts appear to mark intermediate steps in the progression from prokaryotes to eukaryotes.
With regards to an evolutionary understanding, the ubiquitously present proteins can also provide useful information. The Cdc48 proteins VAT, Cpa, and p97 exhibit distinct functional modes in archaea, actinobacteria and eukaryotes, respectively. While VAT and Cpa dock directly onto the CP, the eukaryotic homologues operate upstream of the 26S proteasome, and physical interactions with the 26S have not yet been reported. At this juncture, the in situ as well as ex situ structural characterization of other proteasome-ATPase complexes are necessary. It is important to identify and characterize the entire proteasome interaction network in both eukaryotic and non-eukaryotic degradation pathways, their sequence of interaction and regulatory mechanisms. Such an analysis would provide deeper insights into how proteasomes operate in the different domains of life, and would help to put forward hypotheses regarding their evolutionary connection.
Funding source: Deutsche Forschungsgemeinschaft
Award Identifier / Grant number: SFB-1035/Project A01
Funding statement: This work has been supported by Deutsche Forschungsgemeinschaft SFB-1035/Project A01, Funder Id: http://dx.doi.org/10.13039/501100001659 (to W.B.).
References
Adam, Z., Adamska, I., Nakabayashi, K., Ostersetzer, O., Haussuhl, K., Manuell, A., Zheng, B., Vallon, O., Rodermel, S.R., Shinozaki, K., et al. (2001). Chloroplast and mitochondrial proteases in Arabidopsis. A proposed nomenclature. Plant Physiol. 125, 1912.10.1104/pp.125.4.1912Search in Google Scholar
Albert, S., Schaffer, M., Beck, F., Mosalaganti, S., Asano, S., Thomas, H.F., Plitzko, J.M., Beck, M., Baumeister, W., and Engel, B.D. (2017). Proteasomes tether to two distinct sites at the nuclear pore complex. Proc. Natl. Acad. Sci. USA 114, 13726–13731.10.1073/pnas.1716305114Search in Google Scholar PubMed PubMed Central
Alves dos Santos, C.M., van Kerkhof, P., and Strous, G.J. (2001). The signal transduction of the growth hormone receptor is regulated by the ubiquitin/proteasome system and continues after endocytosis. J. Biol. Chem. 276, 10839–10846.10.1074/jbc.M003635200Search in Google Scholar PubMed
Arendt, C.S. and Hochstrasser, M. (1999). Eukaryotic 20S proteasome catalytic subunit propeptides prevent active site inactivation by N-terminal acetylation and promote particle assembly. EMBO J. 18, 3575–3585.10.1093/emboj/18.13.3575Search in Google Scholar PubMed PubMed Central
Arrigo, A.P., Tanaka, K., Goldberg, A.L., and Welch, W.J. (1988).Identity of the 19S ‘prosome’ particle with the large multifunctional protease complex of mammalian cells (the proteasome). Nature 331, 192–194.10.1038/331192a0Search in Google Scholar PubMed
Asano, S., Fukuda, Y., Beck, F., Aufderheide, A., Forster, F., Danev, R., and Baumeister, W. (2015). Proteasomes. A molecular census of 26S proteasomes in intact neurons. Science 347, 439–442.10.1126/science.1261197Search in Google Scholar PubMed
Aufderheide, A., Beck, F., Stengel, F., Hartwig, M., Schweitzer, A., Pfeifer, G., Goldberg, A.L., Sakata, E., Baumeister, W., and Forster, F. (2015). Structural characterization of the interaction of Ubp6 with the 26S proteasome. Proc. Natl. Acad. Sci. USA 112, 8626–8631.10.1073/pnas.1510449112Search in Google Scholar PubMed PubMed Central
Augustyniak, R. and Kay, L.E. (2018). Cotranslocational processing of the protein substrate calmodulin by an AAA+ unfoldase occurs via unfolding and refolding intermediates. Proc. Natl. Acad. Sci. USA 115, E4786–E4795.10.1073/pnas.1721811115Search in Google Scholar PubMed PubMed Central
Bader, M. and Steller, H. (2009). Regulation of cell death by the ubiquitin-proteasome system. Curr. Opin. Cell Biol. 21, 878–884.10.1016/j.ceb.2009.09.005Search in Google Scholar PubMed PubMed Central
Barthelme, D. and Sauer, R.T. (2012). Identification of the Cdc48*20S proteasome as an ancient AAA+ proteolytic machine. Science 337, 843–846.10.1126/science.1224352Search in Google Scholar PubMed PubMed Central
Barthelme, D., Chen, J.Z., Grabenstatter, J., Baker, T.A., and Sauer, R.T. (2014). Architecture and assembly of the archaeal Cdc48*20S proteasome. Proc. Natl. Acad. Sci. USA 111, E1687–1694.10.1073/pnas.1404823111Search in Google Scholar PubMed PubMed Central
Bashore, C., Dambacher, C.M., Goodall, E.A., Matyskiela, M.E., Lander, G.C., and Martin, A. (2015). Ubp6 deubiquitinase controls conformational dynamics and substrate degradation of the 26S proteasome. Nat. Struct. Mol. Biol. 22, 712–719.10.1038/nsmb.3075Search in Google Scholar
Bauer, M.W., Bauer, S.H., and Kelly, R.M. (1997). Purification and characterization of a proteasome from the Hyperthermophilic Archaeon Pyrococcus furiosus. Appl. Environ. Microbiol. 63, 1160–1164.10.1128/aem.63.3.1160-1164.1997Search in Google Scholar
Baumeister, W., Cejka, Z., Kania, M., and Seemuller, E. (1997). The proteasome: a macromolecular assembly designed to confine proteolysis to a nanocompartment. Biol. Chem. 378, 121–130.Search in Google Scholar
Baumeister, W., Walz, J., Zuhl, F., and Seemuller, E. (1998). The proteasome: paradigm of a self-compartmentalizing protease. Cell 92, 367–380.10.1016/S0092-8674(00)80929-0Search in Google Scholar
Beck, F., Unverdorben, P., Bohn, S., Schweitzer, A., Pfeifer, G., Sakata, E., Nickell, S., Plitzko, J.M., Villa, E., Baumeister, W., et al. (2012). Near-atomic resolution structural model of the yeast 26S proteasome. Proc. Natl. Acad. Sci. USA 109, 14870–14875.10.1073/pnas.1213333109Search in Google Scholar PubMed PubMed Central
Benaroudj, N. and Goldberg, A.L. (2000). PAN, the proteasome-activating nucleotidase from archaebacteria, is a protein-unfolding molecular chaperone. Nat. Cell Biol. 2, 833–839.10.1038/35041081Search in Google Scholar PubMed
Blickwedehl, J., Agarwal, M., Seong, C., Pandita, R.K., Melendy, T., Sung, P., Pandita, T.K., and Bangia, N. (2008). Role for proteasome activator PA200 and postglutamyl proteasome activity in genomic stability. Proc. Natl. Acad. Sci. USA 105, 16165–16170.10.1073/pnas.0803145105Search in Google Scholar PubMed PubMed Central
Bodnar, N. and Rapoport, T. (2017). Toward an understanding of the Cdc48/p97 ATPase. F1000Res. 6, 1318.10.12688/f1000research.11683.1Search in Google Scholar PubMed PubMed Central
Bolten, M., Delley, C.L., Leibundgut, M., Boehringer, D., Ban, N., and Weber-Ban, E. (2016). Structural analysis of the bacterial proteasome activator Bpa in complex with the 20S proteasome. Structure 24, 2138–2151.10.1016/j.str.2016.10.008Search in Google Scholar PubMed
Burns, K.E., Cerda-Maira, F.A., Wang, T., Li, H., Bishai, W.R., and Darwin, K.H. (2010). Depupylation of prokaryotic ubiquitin-like protein from mycobacterial proteasome substrates. Mol. Cell 39, 821–827.10.1016/j.molcel.2010.07.019Search in Google Scholar PubMed PubMed Central
Chen, B., Retzlaff, M., Roos, T., and Frydman, J. (2011). Cellular strategies of protein quality control. Cold Spring Harb. Perspect. Biol. 3, a004374.10.1101/cshperspect.a004374Search in Google Scholar PubMed PubMed Central
Ciechanover, A. (2005). Proteolysis: from the lysosome to ubiquitin and the proteasome. Nat. Rev. Mol. Cell Biol. 6, 79–87.10.1038/nrm1552Search in Google Scholar
Coux, O., Tanaka, K., and Goldberg, A.L. (1996). Structure and functions of the 20S and 26S proteasomes. Annu. Rev. Biochem. 65, 801–847.10.1146/annurev.bi.65.070196.004101Search in Google Scholar
Dahlmann, B. (2007). Role of proteasomes in disease. BMC Biochem. 8 (Suppl 1.), S3.10.1186/1471-2091-8-S1-S3Search in Google Scholar
Dahlmann, B., Kopp, F., Kuehn, L., Niedel, B., Pfeifer, G., Hegerl, R., and Baumeister, W. (1989). The multicatalytic proteinase (prosome) is ubiquitous from eukaryotes to archaebacteria. FEBS Lett. 251, 125–131.10.1016/0014-5793(89)81441-3Search in Google Scholar
Dambacher, C.M., Worden, E.J., Herzik, M.A., Martin, A., and Lander, G.C. (2016). Atomic structure of the 26S proteasome lid reveals the mechanism of deubiquitinase inhibition. eLife 5, e13027.10.7554/eLife.13027Search in Google Scholar
Dange, T., Smith, D., Noy, T., Rommel, P.C., Jurzitza, L., Cordero, R.J., Legendre, A., Finley, D., Goldberg, A.L., and Schmidt, M. (2011). Blm10 protein promotes proteasomal substrate turnover by an active gating mechanism. J. Biol. Chem. 286, 42830–42839.10.1074/jbc.M111.300178Search in Google Scholar
Dantuma, N.P. and Bott, L.C. (2014). The ubiquitin-proteasome system in neurodegenerative diseases: precipitating factor, yet part of the solution. Front Mol. Neurosci. 7, 70.10.3389/fnmol.2014.00070Search in Google Scholar
Darwin, K.H., Lin, G., Chen, Z., Li, H., and Nathan, C.F. (2005). Characterization of a Mycobacterium tuberculosis proteasomal ATPase homologue. Mol. Microbiol. 55, 561–571.10.1111/j.1365-2958.2004.04403.xSearch in Google Scholar
de la Pena, A.H., Goodall, E.A., Gates, S.N., Lander, G.C., and Martin, A. (2018). Substrate-engaged 26S proteasome structures reveal mechanisms for ATP-hydrolysis-driven translocation. Science 362, eaav0725.10.1126/science.aav0725Search in Google Scholar
Deveraux, Q., Ustrell, V., Pickart, C., and Rechsteiner, M. (1994). A 26 S protease subunit that binds ubiquitin conjugates. J. Biol. Chem. 269, 7059–7061.10.1016/S0021-9258(17)37244-7Search in Google Scholar
Dikic, I. (2017). Proteasomal and autophagic degradation systems. Annu. Rev. Biochem. 86, 193–224.10.1146/annurev-biochem-061516-044908Search in Google Scholar PubMed
Ding, Z., Fu, Z., Xu, C., Wang, Y., Wang, Y., Li, J., Kong, L., Chen, J., Li, N., Zhang, R., et al. (2017). High-resolution cryo-EM structure of the proteasome in complex with ADP-AlFx. Cell Res. 27, 373–385.10.1038/cr.2017.12Search in Google Scholar PubMed PubMed Central
Ding, Z., Xu, C., Sahu, I., Wang, Y., Fu, Z., Huang, M., Wong, C.C.L., Glickman, M.H., and Cong, Y. (2019). Structural snapshots of 26S proteasome reveal tetraubiquitin-induced conformations. Mol. Cell 73, 1150–1161.e1156.10.1016/j.molcel.2019.01.018Search in Google Scholar PubMed
Djuranovic, S., Hartmann, M.D., Habeck, M., Ursinus, A., Zwickl, P., Martin, J., Lupas, A.N., and Zeth, K. (2009). Structure and activity of the N-terminal substrate recognition domains in proteasomal ATPases. Mol. Cell 34, 580–590.10.1016/j.molcel.2009.04.030Search in Google Scholar PubMed
Dong, Y., Zhang, S., Wu, Z., Li, X., Wang, W.L., Zhu, Y., Stoilova-McPhie, S., Lu, Y., Finley, D., and Mao, Y. (2019). Cryo-EM structures and dynamics of substrate-engaged human 26S proteasome. Nature 565, 49–55.10.1038/s41586-018-0736-4Search in Google Scholar PubMed PubMed Central
Eisele, M.R., Reed, R.G., Rudack, T., Schweitzer, A., Beck, F., Nagy, I., Pfeifer, G., Plitzko, J.M., Baumeister, W., Tomko Jr, R.J., et al. (2018). Expanded coverage of the 26S proteasome conformational landscape reveals mechanisms of peptidase gating. Cell Rep. 24, 1301–1315 e1305.10.1016/j.celrep.2018.07.004Search in Google Scholar PubMed PubMed Central
Elsasser, S., Gali, R.R., Schwickart, M., Larsen, C.N., Leggett, D.S., Muller, B., Feng, M.T., Tubing, F., Dittmar, G.A., and Finley, D. (2002). Proteasome subunit Rpn1 binds ubiquitin-like protein domains. Nat. Cell Biol. 4, 725–730.10.1038/ncb845Search in Google Scholar PubMed
Fehlker, M., Wendler, P., Lehmann, A., and Enenkel, C. (2003). Blm3 is part of nascent proteasomes and is involved in a late stage of nuclear proteasome assembly. EMBO Rep. 4, 959–963.10.1038/sj.embor.embor938Search in Google Scholar PubMed PubMed Central
Finley, D. (2009). Recognition and processing of ubiquitin-protein conjugates by the proteasome. Annu. Rev. Biochem. 78, 477–513.10.1146/annurev.biochem.78.081507.101607Search in Google Scholar PubMed PubMed Central
Forouzan, D., Ammelburg, M., Hobel, C.F., Stroh, L.J., Sessler, N., Martin, J., and Lupas, A.N. (2012). The archaeal proteasome is regulated by a network of AAA ATPases. J. Biol. Chem. 287, 39254–39262.10.1074/jbc.M112.386458Search in Google Scholar PubMed PubMed Central
Forster, A., Masters, E.I., Whitby, F.G., Robinson, H., and Hill, C.P. (2005). The 1.9 A structure of a proteasome-11S activator complex and implications for proteasome-PAN/PA700 interactions. Mol. Cell 18, 589–599.10.1016/j.molcel.2005.04.016Search in Google Scholar
Fu, X., Liu, R., Sanchez, I., Silva-Sanchez, C., Hepowit, N.L., Cao, S., Chen, S., and Maupin-Furlow, J. (2016). Ubiquitin-like proteasome system represents a eukaryotic-like pathway for targeted proteolysis in Archaea. MBio. 7, e00379–16.10.1128/mBio.00379-16Search in Google Scholar
Fuchs, A.C.D. and Hartmann, M.D. (2019). On the origins of symmetry and modularity in the proteasome family: symmetry transitions are pivotal in the evolution and functional diversification of self-compartmentalizing proteases. Bioessays 41, e1800237.10.1002/bies.201800237Search in Google Scholar
Fuchs, A.C.D., Alva, V., Maldoner, L., Albrecht, R., Hartmann, M.D., and Martin, J. (2017). The architecture of the Anbu complex reflects an evolutionary intermediate at the origin of the proteasome system. Structure 25, 834–845 e835.10.1016/j.str.2017.04.005Search in Google Scholar
Fuchs, A.C.D., Maldoner, L., Hipp, K., Hartmann, M.D., and Martin, J. (2018). Structural characterization of the bacterial proteasome homolog BPH reveals a tetradecameric double-ring complex with unique inner cavity properties. J. Biol. Chem. 293, 920–930.10.1074/jbc.M117.815258Search in Google Scholar
Gerega, A., Rockel, B., Peters, J., Tamura, T., Baumeister, W., and Zwickl, P. (2005). VAT, the Thermoplasma homolog of mammalian p97/VCP, is an N domain-regulated protein unfoldase. J. Biol. Chem. 280, 42856–42862.10.1074/jbc.M510592200Search in Google Scholar
Glickman, M.H., Rubin, D.M., Coux, O., Wefes, I., Pfeifer, G., Cjeka, Z., Baumeister, W., Fried, V.A., and Finley, D. (1998). A subcomplex of the proteasome regulatory particle required for ubiquitin-conjugate degradation and related to the COP9-signalosome and eIF3. Cell 94, 615–623.10.1016/S0092-8674(00)81603-7Search in Google Scholar
Golbik, R., Lupas, A.N., Koretke, K.K., Baumeister, W., and Peters, J. (1999). The Janus face of the archaeal Cdc48/p97 homologue VAT: protein folding versus unfolding. Biol. Chem. 380, 1049–1062.10.1515/BC.1999.131Search in Google Scholar PubMed
Goldberg, A.L. (2007). Functions of the proteasome: from protein degradation and immune surveillance to cancer therapy. Biochem. Soc. Trans. 35, 12–17.10.1042/BST0350012Search in Google Scholar PubMed
Goldberg, A.L. (2012). Development of proteasome inhibitors as research tools and cancer drugs. J. Cell. Biol. 199, 583–588.10.1083/jcb.201210077Search in Google Scholar PubMed PubMed Central
Gomez, T.A., Kolawa, N., Gee, M., Sweredoski, M.J., and Deshaies, R.J. (2011). Identification of a functional docking site in the Rpn1 LRR domain for the UBA-UBL domain protein Ddi1. BMC Biol. 9, 33.10.1186/1741-7007-9-33Search in Google Scholar
Groettrup, M., Kirk, C.J., and Basler, M. (2010). Proteasomes in immune cells: more than peptide producers? Nat. Rev. Immunol. 10, 73–78.10.1038/nri2687Search in Google Scholar
Groll, M. and Huber, R. (2003). Substrate access and processing by the 20S proteasome core particle. Int. J. Biochem. Cell Biol. 35, 606–616.10.1016/S1357-2725(02)00390-4Search in Google Scholar
Groll, M., Heinemeyer, W., Jager, S., Ullrich, T., Bochtler, M., Wolf, D.H., and Huber, R. (1999). The catalytic sites of 20S proteasomes and their role in subunit maturation: a mutational and crystallographic study. Proc. Natl. Acad. Sci. USA 96, 10976–10983.10.1073/pnas.96.20.10976Search in Google Scholar
Guo, Q., Lehmer, C., Martinez-Sanchez, A., Rudack, T., Beck, F., Hartmann, H., Perez-Berlanga, M., Frottin, F., Hipp, M.S., Hartl, F.U., et al. (2018). In situ structure of neuronal C9orf72 Poly-GA aggregates reveals proteasome recruitment. Cell 172, 696–705, e612.10.1016/j.cell.2017.12.030Search in Google Scholar
Gur, E., Biran, D., and Ron, E.Z. (2011). Regulated proteolysis in Gram-negative bacteria – how and when? Nat. Rev. Microbiol. 9, 839–848.10.1038/nrmicro2669Search in Google Scholar
Hamazaki, J., Iemura, S., Natsume, T., Yashiroda, H., Tanaka, K., and Murata, S. (2006). A novel proteasome interacting protein recruits the deubiquitinating enzyme UCH37 to 26S proteasomes. EMBO J. 25, 4524–4536.10.1038/sj.emboj.7601338Search in Google Scholar
Harris, J.R. (1968). Release of a macromolecular protein component from human erythrocyte ghosts. Biochim. Biophys. Acta. 150, 534–537.10.1016/0005-2736(68)90157-0Search in Google Scholar
He, J., Cui, L., Zeng, Y., Wang, G., Zhou, P., Yang, Y., Ji, L., Zhao, Y., Chen, J., Wang, Z., et al. (2012). REGgamma is associated with multiple oncogenic pathways in human cancers. BMC Cancer 12, 75.10.1186/1471-2407-12-75Search in Google Scholar
Hegerl, R., Pfeifer, G., Pühler, G., Dahlmann, B., and Baumeister, W. (1991). The three-dimensional structure of proteasomes from Thermoplasma acidophilum as determined by electron microscopy using random conical tilting. FEBS Lett. 283, 117–121.10.1016/0014-5793(91)80567-MSearch in Google Scholar
Heinemeyer, W., Fischer, M., Krimmer, T., Stachon, U., and Wolf, D.H. (1997). The active sites of the eukaryotic 20 S proteasome and their involvement in subunit precursor processing. J. Biol. Chem. 272, 25200–25209.10.1074/jbc.272.40.25200Search in Google Scholar PubMed
Hendil, K.B., Khan, S., and Tanaka, K. (1998). Simultaneous binding of PA28 and PA700 activators to 20 S proteasomes. Biochem. J. 332, 749–754.10.1042/bj3320749Search in Google Scholar PubMed PubMed Central
Herzik Jr, M.A., Wu, M., and Lander, G.C. (2017). Achieving better-than-3-Å resolution by single-particle cryo-EM at 200 keV. Nat. Methods 14, 1075–1078.10.1038/nmeth.4461Search in Google Scholar PubMed PubMed Central
Hu, K., Jastrab, J.B., Zhang, S., Kovach, A., Zhao, G., Darwin, K.H., and Li, H. (2018). Proteasome substrate capture and gate opening by the accessory factor PafE from Mycobacterium tuberculosis. J. Biol. Chem. 293, 4713–4723.10.1074/jbc.RA117.001471Search in Google Scholar PubMed PubMed Central
Huang, R., Ripstein, Z.A., Augustyniak, R., Lazniewski, M., Ginalski, K., Kay, L.E., and Rubinstein, J.L. (2016). Unfolding the mechanism of the AAA+ unfoldase VAT by a combined cryo-EM, solution NMR study. Proc. Natl. Acad. Sci. USA 113, E4190–4199.10.1073/pnas.1603980113Search in Google Scholar PubMed PubMed Central
Huber, E.M., Heinemeyer, W., Li, X., Arendt, C.S., Hochstrasser, M., and Groll, M. (2016). A unified mechanism for proteolysis and autocatalytic activation in the 20S proteasome. Nat. Commun. 7, 10900.10.1038/ncomms10900Search in Google Scholar PubMed PubMed Central
Humbard, M.A., Miranda, H.V., Lim, J.M., Krause, D.J., Pritz, J.R., Zhou, G., Chen, S., Wells, L., and Maupin-Furlow, J.A. (2010). Ubiquitin-like small archaeal modifier proteins (SAMPs) in Haloferax volcanii. Nature 463, 54–60.10.1038/nature08659Search in Google Scholar PubMed PubMed Central
Husnjak, K., Elsasser, S., Zhang, N., Chen, X., Randles, L., Shi, Y., Hofmann, K., Walters, K.J., Finley, D., and Dikic, I. (2008). Proteasome subunit Rpn13 is a novel ubiquitin receptor. Nature 453, 481–488.10.1038/nature06926Search in Google Scholar PubMed PubMed Central
Isasa, M., Katz, E.J., Kim, W., Yugo, V., Gonzalez, S., Kirkpatrick, D.S., Thomson, T.M., Finley, D., Gygi, S.P., and Crosas, B. (2010). Monoubiquitination of RPN10 regulates substrate recruitment to the proteasome. Mol. Cell 38, 733–745.10.1016/j.molcel.2010.05.001Search in Google Scholar PubMed PubMed Central
Jager, S., Groll, M., Huber, R., Wolf, D.H., and Heinemeyer, W. (1999). Proteasome beta-type subunits: unequal roles of propeptides in core particle maturation and a hierarchy of active site function. J. Mol. Biol. 291, 997–1013.10.1006/jmbi.1999.2995Search in Google Scholar PubMed
Jastrab, J.B., Wang, T., Murphy, J.P., Bai, L., Hu, K., Merkx, R., Huang, J., Chatterjee, C., Ovaa, H., Gygi, S.P., et al. (2015). An adenosine triphosphate-independent proteasome activator contributes to the virulence of Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 112, E1763–1772.10.1073/pnas.1423319112Search in Google Scholar PubMed PubMed Central
Jiao, L., Ouyang, S., Shaw, N., Song, G., Feng, Y., Niu, F., Qiu, W., Zhu, H., Hung, L.W., Zuo, X., et al. (2014). Mechanism of the Rpn13-induced activation of Uch37. Protein Cell 5, 616–630.10.1007/s13238-014-0046-zSearch in Google Scholar PubMed PubMed Central
Khor, B., Bredemeyer, A.L., Huang, C.Y., Turnbull, I.R., Evans, R., Maggi Jr, L.B., White, J.M., Walker, L.M., Carnes, K., Hess, R.A., et al. (2006). Proteasome activator PA200 is required for normal spermatogenesis. Mol. Cell Biol. 26, 2999–3007.10.1128/MCB.26.8.2999-3007.2006Search in Google Scholar PubMed PubMed Central
Kim, H.T. and Goldberg, A.L. (2017). The deubiquitinating enzyme Usp14 allosterically inhibits multiple proteasomal activities and ubiquitin-independent proteolysis. J. Biol. Chem. 292, 9830–9839.10.1074/jbc.M116.763128Search in Google Scholar PubMed PubMed Central
Kim, P.K., Hailey, D.W., Mullen, R.T., and Lippincott-Schwartz, J. (2008). Ubiquitin signals autophagic degradation of cytosolic proteins and peroxisomes. Proc. Natl. Acad. Sci. USA 105, 20567–20574.10.1073/pnas.0810611105Search in Google Scholar PubMed PubMed Central
Kim, Y.C., Snoberger, A., Schupp, J., and Smith, D.M. (2015). ATP binding to neighbouring subunits and intersubunit allosteric coupling underlie proteasomal ATPase function. Nat. Commun. 6, 8520.10.1038/ncomms9520Search in Google Scholar PubMed PubMed Central
Knipfer, N. and Shrader, T.E. (1997). Inactivation of the 20S proteasome in Mycobacterium smegmatis. Mol. Microbiol. 25, 375–383.10.1046/j.1365-2958.1997.4721837.xSearch in Google Scholar PubMed
Koepp, D.M. (2014). Cell cycle regulation by protein degradation. Methods Mol. Biol. 1170, 61–73.10.1007/978-1-4939-0888-2_4Search in Google Scholar PubMed
Komander, D. and Rape, M. (2012). The ubiquitin code. Annu. Rev. Biochem. 81, 203–229.10.1146/annurev-biochem-060310-170328Search in Google Scholar PubMed
Korolchuk, V.I., Menzies, F.M., and Rubinsztein, D.C. (2010). Mechanisms of cross-talk between the ubiquitin-proteasome and autophagy-lysosome systems. FEBS Lett. 584, 1393–1398.10.1016/j.febslet.2009.12.047Search in Google Scholar PubMed
Kumoi, K., Satoh, T., Murata, K., Hiromoto, T., Mizushima, T., Kamiya, Y., Noda, M., Uchiyama, S., Yagi, H., and Kato, K. (2013). An archaeal homolog of proteasome assembly factor functions as a proteasome activator. PLoS One 8, e60294.10.1371/journal.pone.0060294Search in Google Scholar
Kunjappu, M.J. and Hochstrasser, M. (2014). Assembly of the 20S proteasome. Biochim. Biophys. Acta 1843, 2–12.10.1016/j.bbamcr.2013.03.008Search in Google Scholar
Kwon, Y.D., Nagy, I., Adams, P.D., Baumeister, W., and Jap, B.K. (2004). Crystal structures of the Rhodococcus proteasome with and without its pro-peptides: implications for the role of the pro-peptide in proteasome assembly. J. Mol. Biol. 335, 233–245.10.1016/j.jmb.2003.08.029Search in Google Scholar
Lam, Y.A., Lawson, T.G., Velayutham, M., Zweier, J.L., and Pickart, C.M. (2002). A proteasomal ATPase subunit recognizes the polyubiquitin degradation signal. Nature 416, 763–767.10.1038/416763aSearch in Google Scholar
Lander, G.C., Estrin, E., Matyskiela, M.E., Bashore, C., Nogales, E., and Martin, A. (2012). Complete subunit architecture of the proteasome regulatory particle. Nature 482, 186–191.10.1038/nature10774Search in Google Scholar
Lasker, K., Forster, F., Bohn, S., Walzthoeni, T., Villa, E., Unverdorben, P., Beck, F., Aebersold, R., Sali, A., and Baumeister, W. (2012). Molecular architecture of the 26S proteasome holocomplex determined by an integrative approach. Proc. Natl. Acad. Sci. USA 109, 1380–1387.10.1073/pnas.1120559109Search in Google Scholar
Leggett, D.S., Hanna, J., Borodovsky, A., Crosas, B., Schmidt, M., Baker, R.T., Walz, T., Ploegh, H., and Finley, D. (2002). Multiple associated proteins regulate proteasome structure and function. Mol. Cell 10, 495–507.10.1016/S1097-2765(02)00638-XSearch in Google Scholar
Lehmann, A., Jechow, K., and Enenkel, C. (2008). Blm10 binds to pre-activated proteasome core particles with open gate conformation. EMBO Rep. 9, 1237–1243.10.1038/embor.2008.190Search in Google Scholar PubMed PubMed Central
Liu, C.W., Corboy, M.J., DeMartino, G.N., and Thomas, P.J. (2003). Endoproteolytic activity of the proteasome. Science 299, 408–411.10.1126/science.1079293Search in Google Scholar PubMed PubMed Central
Lu, X., Nowicka, U., Sridharan, V., Liu, F., Randles, L., Hymel, D., Dyba, M., Tarasov, S.G., Tarasova, N.I., Zhao, X.Z., et al. (2017). Structure of the Rpn13-Rpn2 complex provides insights for Rpn13 and Uch37 as anticancer targets. Nat. Commun. 8, 15540.10.1038/ncomms15540Search in Google Scholar PubMed PubMed Central
Lupoli, T.J., Vaubourgeix, J., Burns-Huang, K., and Gold, B. (2018). Targeting the proteostasis network for mycobacterial drug discovery. ACS Infect Dis. 4, 478–498.10.1021/acsinfecdis.7b00231Search in Google Scholar
Majumder, P., Rudack, T., Beck, F., Danev, R., Pfeifer, G., Nagy, I., and Baumeister, W. (2019). Cryo-EM structures of the archaeal PAN-proteasome reveal an around-the-ring ATPase cycle. Proc. Natl. Acad. Sci. U.S.A. 116, 534–539.10.1073/pnas.1817752116Search in Google Scholar
Martin, A., Baker, T.A., and Sauer, R.T. (2007). Distinct static and dynamic interactions control ATPase-peptidase communication in a AAA+ protease. Mol. Cell 27, 41–52.10.1016/j.molcel.2007.05.024Search in Google Scholar
Martin, A., Baker, T.A., and Sauer, R.T. (2008). Pore loops of the AAA+ ClpX machine grip substrates to drive translocation and unfolding. Nat. Struct. Mol. Biol. 15, 1147–1151.10.1038/nsmb.1503Search in Google Scholar
Masson, P., Lundgren, J., and Young, P. (2003). Drosophila proteasome regulator REGγ: transcriptional activation by DNA replication-related factor DREF and evidence for a role in cell cycle progression. J. Mol. Biol. 327, 1001–1012.10.1016/S0022-2836(03)00188-8Search in Google Scholar
Matiuhin, Y., Kirkpatrick, D.S., Ziv, I., Kim, W., Dakshinamurthy, A., Kleifeld, O., Gygi, S.P., Reis, N., and Glickman, M.H. (2008). Extraproteasomal Rpn10 restricts access of the polyubiquitin-binding protein Dsk2 to proteasome. Mol. Cell 32, 415–425.10.1016/j.molcel.2008.10.011Search in Google Scholar PubMed PubMed Central
Matyskiela, M.E., Lander, G.C., and Martin, A. (2013). Conformational switching of the 26S proteasome enables substrate degradation. Nat. Struct. Mol. Biol. 20, 781–788.10.1038/nsmb.2616Search in Google Scholar PubMed PubMed Central
Maupin-Furlow, J.A. (2018). Proteolytic systems of archaea: slicing, dicing, and mincing in the extreme. Emerging Topics Life Sci., ETLS20180025.10.1042/ETLS20180025Search in Google Scholar
Maupin-Furlow, J.A., Aldrich, H.C., and Ferry, J.G. (1998). Biochemical characterization of the 20S proteasome from the methanoarchaeon Methanosarcina thermophila. J. Bacteriol. 180, 1480–1487.10.1128/JB.180.6.1480-1487.1998Search in Google Scholar PubMed PubMed Central
Meyer, H., Bug, M., and Bremer, S. (2012). Emerging functions of the VCP/p97 AAA-ATPase in the ubiquitin system. Nat. Cell Biol. 14, 117–123.10.1038/ncb2407Search in Google Scholar PubMed
Miller, J.M. and Enemark, E.J. (2016). Fundamental characteristics of AAA+ protein family structure and function. Archaea 2016, 9294307.10.1155/2016/9294307Search in Google Scholar PubMed PubMed Central
Motegi, A., Murakawa, Y., and Takeda, S. (2009). The vital link between the ubiquitin-proteasome pathway and DNA repair: impact on cancer therapy. Cancer Lett. 283, 1–9.10.1016/j.canlet.2008.12.030Search in Google Scholar
Müller, A.U. and Weber-Ban, E. (2019). The bacterial proteasome at the core of diverse degradation pathways. Front Mol. Biosci. 6, 23.10.3389/fmolb.2019.00023Search in Google Scholar
Murata, S., Kawahara, H., Tohma, S., Yamamoto, K., Kasahara, M., Nabeshima, Y., Tanaka, K., and Chiba, T. (1999). Growth retardation in mice lacking the proteasome activator PA28γ. J. Biol. Chem. 274, 38211–38215.10.1074/jbc.274.53.38211Search in Google Scholar
Murata, S., Yashiroda, H., and Tanaka, K. (2009). Molecular mechanisms of proteasome assembly. Nat. Rev. Mol. Cell Biol. 10, 104–115.10.1038/nrm2630Search in Google Scholar
Murata, S., Takahama, Y., Kasahara, M., and Tanaka, K. (2018). The immunoproteasome and thymoproteasome: functions, evolution and human disease. Nat. Immunol. 19, 923–931.10.1038/s41590-018-0186-zSearch in Google Scholar
Navon, A. and Goldberg, A.L. (2001). Proteins are unfolded on the surface of the ATPase ring before transport into the proteasome. Mol. Cell 8, 1339–1349.10.1016/S1097-2765(01)00407-5Search in Google Scholar
Ogura, T., Whiteheart, S.W., and Wilkinson, A.J. (2004). Conserved arginine residues implicated in ATP hydrolysis, nucleotide-sensing, and inter-subunit interactions in AAA and AAA+ATPases. J. Struct. Biol. 146, 106–112.10.1016/j.jsb.2003.11.008Search in Google Scholar
Orlowski, M. (1990). The multicatalytic proteinase complex, a major extralysosomal proteolytic system. Biochemistry 29, 10289–10297.10.1021/bi00497a001Search in Google Scholar
Pamnani, V., Tamura, T., Lupas, A., Peters, J., Cejka, Z., Ashraf, W., and Baumeister, W. (1997). Cloning, sequencing and expression of VAT, a CDC48/p97 ATPase homologue from the archaeon Thermoplasma acidophilum. FEBS Lett. 404, 263–268.10.1016/S0014-5793(97)00138-5Search in Google Scholar
Paraskevopoulos, K., Kriegenburg, F., Tatham, M.H., Rosner, H.I., Medina, B., Larsen, I.B., Brandstrup, R., Hardwick, K.G., Hay, R.T., Kragelund, B.B., et al. (2014). Dss1 is a 26S proteasome ubiquitin receptor. Mol. Cell 56, 453–461.10.1016/j.molcel.2014.09.008Search in Google Scholar PubMed PubMed Central
Pathare, G.R., Nagy, I., Bohn, S., Unverdorben, P., Hubert, A., Korner, R., Nickell, S., Lasker, K., Sali, A., Tamura, T., et al. (2012). The proteasomal subunit Rpn6 is a molecular clamp holding the core and regulatory subcomplexes together. Proc. Natl. Acad. Sci. USA 109, 149–154.10.1073/pnas.1117648108Search in Google Scholar PubMed PubMed Central
Pathare, G.R., Nagy, I., Sledz, P., Anderson, D.J., Zhou, H.J., Pardon, E., Steyaert, J., Forster, F., Bracher, A., and Baumeister, W. (2014). Crystal structure of the proteasomal deubiquitylation module Rpn8-Rpn11. Proc. Natl. Acad. Sci. USA 111, 2984–2989.10.1073/pnas.1400546111Search in Google Scholar PubMed PubMed Central
Pearce, M.J., Mintseris, J., Ferreyra, J., Gygi, S.P., and Darwin, K.H. (2008). Ubiquitin-like protein involved in the proteasome pathway of Mycobacterium tuberculosis. Science 322, 1104–1107.10.1126/science.1163885Search in Google Scholar PubMed PubMed Central
Rabl, J., Smith, D.M., Yu, Y., Chang, S.C., Goldberg, A.L., and Cheng, Y. (2008). Mechanism of gate opening in the 20S proteasome by the proteasomal ATPases. Mol. Cell 30, 360–368.10.1016/j.molcel.2008.03.004Search in Google Scholar PubMed PubMed Central
Ravid, T. and Hochstrasser, M. (2008). Diversity of degradation signals in the ubiquitin-proteasome system. Nat. Rev. Mol. Cell Biol. 9, 679–690.10.1038/nrm2468Search in Google Scholar PubMed PubMed Central
Ripstein, Z.A., Huang, R., Augustyniak, R., Kay, L.E., and Rubinstein, J.L. (2017). Structure of a AAA+ unfoldase in the process of unfolding substrate. eLife 6, e25754.10.7554/eLife.25754Search in Google Scholar PubMed PubMed Central
Rohrwild, M., Coux, O., Huang, H.C., Moerschell, R.P., Yoo, S.J., Seol, J.H., Chung, C.H., and Goldberg, A.L. (1996). HslV-HslU: a novel ATP-dependent protease complex in Escherichia coli related to the eukaryotic proteasome. Proc. Natl. Acad. Sci. U.S.A. 93, 5808–5813.10.1073/pnas.93.12.5808Search in Google Scholar PubMed PubMed Central
Rosenzweig, R., Bronner, V., Zhang, D., Fushman, D., and Glickman, M.H. (2012). Rpn1 and Rpn2 coordinate ubiquitin processing factors at proteasome. J. Biol. Chem. 287, 14659–14671.10.1074/jbc.M111.316323Search in Google Scholar PubMed PubMed Central
Ruschak, A.M., Religa, T.L., Breuer, S., Witt, S., and Kay, L.E. (2010). The proteasome antechamber maintains substrates in an unfolded state. Nature 467, 868–871.10.1038/nature09444Search in Google Scholar PubMed
Sadre-Bazzaz, K., Whitby, F.G., Robinson, H., Formosa, T., and Hill, C.P. (2010). Structure of a Blm10 complex reveals common mechanisms for proteasome binding and gate opening. Mol. Cell 37, 728–735.10.1016/j.molcel.2010.02.002Search in Google Scholar PubMed PubMed Central
Samanovic, M.I. and Darwin, K.H. (2016). Game of ’Somes: protein destruction for Mycobacterium tuberculosis pathogenesis. Trends Microbiol. 24, 26–34.10.1016/j.tim.2015.10.001Search in Google Scholar PubMed PubMed Central
Schmidt, M. and Finley, D. (2014). Regulation of proteasome activity in health and disease. Biochim. Biophys. Acta 1843, 13–25.10.1016/j.bbamcr.2013.08.012Search in Google Scholar PubMed PubMed Central
Schmidt, M., Haas, W., Crosas, B., Santamaria, P.G., Gygi, S.P., Walz, T., and Finley, D. (2005). The HEAT repeat protein Blm10 regulates the yeast proteasome by capping the core particle. Nat. Struct. Mol. Biol. 12, 294–303.10.1038/nsmb914Search in Google Scholar PubMed
Schweitzer, A., Aufderheide, A., Rudack, T., Beck, F., Pfeifer, G., Plitzko, J.M., Sakata, E., Schulten, K., Forster, F., and Baumeister, W. (2016). Structure of the human 26S proteasome at a resolution of 3.9 Å. Proc. Natl. Acad. Sci. USA 113, 7816–7821.10.1073/pnas.1608050113Search in Google Scholar PubMed PubMed Central
Seemuller, E., Lupas, A., Stock, D., Lowe, J., Huber, R., and Baumeister, W. (1995). Proteasome from Thermoplasma acidophilum: a threonine protease. Science 268, 579–582.10.1126/science.7725107Search in Google Scholar PubMed
Seemuller, E., Lupas, A., and Baumeister, W. (1996). Autocatalytic processing of the 20S proteasome. Nature 382, 468–471.10.1038/382468a0Search in Google Scholar PubMed
Sharon, M., Witt, S., Felderer, K., Rockel, B., Baumeister, W., and Robinson, C.V. (2006). 20S proteasomes have the potential to keep substrates in store for continual degradation. J. Biol. Chem. 281, 9569–9575.10.1074/jbc.M511951200Search in Google Scholar PubMed
Shi, Y., Chen, X., Elsasser, S., Stocks, B.B., Tian, G., Lee, B.H., Shi, Y., Zhang, N., de Poot, S.A., Tuebing, F., et al. (2016). Rpn1 provides adjacent receptor sites for substrate binding and deubiquitination by the proteasome. Science 351, aad9421.10.1126/science.aad9421Search in Google Scholar PubMed PubMed Central
Sledz, P., Unverdorben, P., Beck, F., Pfeifer, G., Schweitzer, A., Forster, F., and Baumeister, W. (2013). Structure of the 26S proteasome with ATP-gammaS bound provides insights into the mechanism of nucleotide-dependent substrate translocation. Proc. Natl. Acad. Sci. USA 110, 7264–7269.10.1073/pnas.1305782110Search in Google Scholar PubMed PubMed Central
Smith, D.M., Kafri, G., Cheng, Y., Ng, D., Walz, T., and Goldberg, A.L. (2005). ATP binding to PAN or the 26S ATPases causes association with the 20S proteasome, gate opening, and translocation of unfolded proteins. Mol. Cell 20, 687–698.10.1016/j.molcel.2005.10.019Search in Google Scholar PubMed
Smith, D.M., Chang, S.C., Park, S., Finley, D., Cheng, Y., and Goldberg, A.L. (2007). Docking of the proteasomal ATPases’ carboxyl termini in the 20S proteasome’s alpha ring opens the gate for substrate entry. Mol. Cell 27, 731–744.10.1016/j.molcel.2007.06.033Search in Google Scholar
Smith, D.M., Fraga, H., Reis, C., Kafri, G., and Goldberg, A.L. (2011). ATP binds to proteasomal ATPases in pairs with distinct functional effects, implying an ordered reaction cycle. Cell 144, 526–538.10.1016/j.cell.2011.02.005Search in Google Scholar
Snoberger, A., Brettrager, E.J., and Smith, D.M. (2018). Conformational switching in the coiled-coil domains of a proteasomal ATPase regulates substrate processing. Nat. Commun. 9, 2374.10.1038/s41467-018-04731-6Search in Google Scholar
Stadtmueller, B.M. and Hill, C.P. (2011). Proteasome activators. Mol. Cell 41, 8–19.10.1016/j.molcel.2010.12.020Search in Google Scholar
Striebel, F., Imkamp, F., Sutter, M., Steiner, M., Mamedov, A., and Weber-Ban, E. (2009). Bacterial ubiquitin-like modifier Pup is deamidated and conjugated to substrates by distinct but homologous enzymes. Nat. Struct. Mol. Biol. 16, 647–651.10.1038/nsmb.1597Search in Google Scholar
Striebel, F., Hunkeler, M., Summer, H., and Weber-Ban, E. (2010). The mycobacterial Mpa-proteasome unfolds and degrades pupylated substrates by engaging Pup’s N-terminus. EMBO J. 29, 1262–1271.10.1038/emboj.2010.23Search in Google Scholar
Sutter, M., Striebel, F., Damberger, F.F., Allain, F.H., and Weber-Ban, E. (2009). A distinct structural region of the prokaryotic ubiquitin-like protein (Pup) is recognized by the N-terminal domain of the proteasomal ATPase Mpa. FEBS Lett. 583, 3151–3157.10.1016/j.febslet.2009.09.020Search in Google Scholar
Tamura, T., Nagy, I., Lupas, A., Lottspeich, F., Cejka, Z., Schoofs, G., Tanaka, K., De Mot, R., and Baumeister, W. (1995). The first characterization of a eubacterial proteasome: the 20S complex of Rhodococcus. Curr. Biol. 5, 766–774.10.1016/S0960-9822(95)00153-9Search in Google Scholar
Tomko Jr, R.J., Funakoshi, M., Schneider, K., Wang, J., and Hochstrasser, M. (2010). Heterohexameric ring arrangement of the eukaryotic proteasomal ATPases: implications for proteasome structure and assembly. Mol. Cell 38, 393–403.10.1016/j.molcel.2010.02.035Search in Google Scholar PubMed PubMed Central
Toste Rego, A. and da Fonseca, P.C.A. (2019). Characterization of fully recombinant human 20S and 20S-PA200 proteasome complexes. Mol. Cell 76, 138–147.10.1016/j.molcel.2019.07.014Search in Google Scholar PubMed PubMed Central
Twomey, E.C., Ji, Z., Wales, T.E., Bodnar, N.O., Ficarro, S.B., Marto, J.A., Engen, J.R., and Rapoport, T.A. (2019). Substrate processing by the Cdc48 ATPase complex is initiated by ubiquitin unfolding. Science 365, eaax1033.10.1126/science.aax1033Search in Google Scholar PubMed PubMed Central
Ustrell, V., Hoffman, L., Pratt, G., and Rechsteiner, M. (2002). PA200, a nuclear proteasome activator involved in DNA repair. EMBO J. 21, 3516–3525.10.1093/emboj/cdf333Search in Google Scholar PubMed PubMed Central
Vander Linden, R.T., Hemmis, C.W., Schmitt, B., Ndoja, A., Whitby, F.G., Robinson, H., Cohen, R.E., Yao, T., and Hill, C.P. (2015). Structural basis for the activation and inhibition of the UCH37 deubiquitylase. Mol. Cell 57, 901–911.10.1016/j.molcel.2015.01.016Search in Google Scholar PubMed PubMed Central
Verma, R., Oania, R., Graumann, J., and Deshaies, R.J. (2004). Multiubiquitin chain receptors define a layer of substrate selectivity in the ubiquitin-proteasome system. Cell 118, 99–110.10.1016/j.cell.2004.06.014Search in Google Scholar PubMed
Wang, Q., Young, P., and Walters, K.J. (2005). Structure of S5a bound to monoubiquitin provides a model for polyubiquitin recognition. J. Mol. Biol. 348, 727–739.10.1016/j.jmb.2005.03.007Search in Google Scholar PubMed
Wang, T., Li, H., Lin, G., Tang, C., Li, D., Nathan, C., Darwin, K.H., and Li, H. (2009). Structural insights on the Mycobacterium tuberculosis proteasomal ATPase Mpa. Structure 17, 1377–1385.10.1016/j.str.2009.08.010Search in Google Scholar PubMed PubMed Central
Wang, T., Darwin, K.H., and Li, H. (2010). Binding-induced folding of prokaryotic ubiquitin-like protein on the Mycobacterium proteasomal ATPase targets substrates for degradation. Nat. Struct. Mol. Biol. 17, 1352–1357.10.1038/nsmb.1918Search in Google Scholar PubMed PubMed Central
Wehmer, M., Rudack, T., Beck, F., Aufderheide, A., Pfeifer, G., Plitzko, J.M., Forster, F., Schulten, K., Baumeister, W., and Sakata, E. (2017). Structural insights into the functional cycle of the ATPase module of the 26S proteasome. Proc. Natl. Acad. Sci. USA 114, 1305–1310.10.1073/pnas.1621129114Search in Google Scholar PubMed PubMed Central
Wendler, P., Ciniawsky, S., Kock, M., and Kube, S. (2012). Structure and function of the AAA+ nucleotide binding pocket. Biochim. Biophys. Acta 1823, 2–14.10.1016/j.bbamcr.2011.06.014Search in Google Scholar PubMed
Wenzel, T. and Baumeister, W. (1995). Conformational constraints in protein degradation by the 20S proteasome. Nat. Struct. Biol. 2, 199–204.10.1038/nsb0395-199Search in Google Scholar PubMed
Wilk, S. and Orlowski, M. (1980). Cation-sensitive neutral endopeptidase: isolation and specificity of the bovine pituitary enzyme. J. Neurochem. 35, 1172–1182.10.1111/j.1471-4159.1980.tb07873.xSearch in Google Scholar PubMed
Wolf, S., Nagy, I., Lupas, A., Pfeifer, G., Cejka, Z., Muller, S.A., Engel, A., De Mot, R., and Baumeister, W. (1998). Characterization of ARC, a divergent member of the AAA ATPase family from Rhodococcus erythropolis. J. Mol. Biol. 277, 13–25.10.1006/jmbi.1997.1589Search in Google Scholar PubMed
Worden, E.J., Padovani, C., and Martin, A. (2014). Structure of the Rpn11-Rpn8 dimer reveals mechanisms of substrate deubiquitination during proteasomal degradation. Nat. Struct. Mol. Biol. 21, 220–227.10.1038/nsmb.2771Search in Google Scholar PubMed
Wu, X. and Rapoport, T.A. (2018). Mechanistic insights into ER-associated protein degradation. Curr. Opin. Cell Biol. 53, 22–28.10.1016/j.ceb.2018.04.004Search in Google Scholar PubMed PubMed Central
Wu, Y., Hu, K., Li, D., Bai, L., Yang, S., Jastrab, J.B., Xiao, S., Hu, Y., Zhang, S., Darwin, K.H., et al. (2017). Mycobacterium tuberculosis proteasomal ATPase Mpa has a β-grasp domain that hinders docking with the proteasome core protease. Mol. Microbiol. 105, 227–241.10.1111/mmi.13695Search in Google Scholar PubMed PubMed Central
Yao, T., Song, L., Xu, W., DeMartino, G.N., Florens, L., Swanson, S.K., Washburn, M.P., Conaway, R.C., Conaway, J.W., and Cohen, R.E. (2006). Proteasome recruitment and activation of the Uch37 deubiquitinating enzyme by Adrm1. Nat. Cell Biol. 8, 994–1002.10.1038/ncb1460Search in Google Scholar PubMed
Yedidi, R.S., Wendler, P., and Enenkel, C. (2017). AAA-ATPases in protein degradation. Front Mol. Biosci. 4, 42.10.3389/fmolb.2017.00042Search in Google Scholar PubMed PubMed Central
Zhang, F., Hu, M., Tian, G., Zhang, P., Finley, D., Jeffrey, P.D., and Shi, Y. (2009a). Structural insights into the regulatory particle of the proteasome from Methanocaldococcus jannaschii. Mol. Cell 34, 473–484.10.1016/j.molcel.2009.04.021Search in Google Scholar PubMed PubMed Central
Zhang, F., Wu, Z., Zhang, P., Tian, G., Finley, D., and Shi, Y. (2009b). Mechanism of substrate unfolding and translocation by the regulatory particle of the proteasome from Methanocaldococcus jannaschii. Mol. Cell 34, 485–496.10.1016/j.molcel.2009.04.022Search in Google Scholar PubMed PubMed Central
Zhou, G., Kowalczyk, D., Humbard, M.A., Rohatgi, S., and Maupin-Furlow, J.A. (2008). Proteasomal components required for cell growth and stress responses in the haloarchaeon Haloferax volcanii. J. Bacteriol. 190, 8096–8105.10.1128/JB.01180-08Search in Google Scholar PubMed PubMed Central
Zhu, Y., Wang, W.L., Yu, D., Ouyang, Q., Lu, Y., and Mao, Y. (2018). Structural mechanism for nucleotide-driven remodeling of the AAA-ATPase unfoldase in the activated human 26S proteasome. Nat. Commun. 9, 1360.10.1038/s41467-018-03785-wSearch in Google Scholar
Ziemski, M., Jomaa, A., Mayer, D., Rutz, S., Giese, C., Veprintsev, D., and Weber-Ban, E. (2018). Cdc48-like protein of actinobacteria (Cpa) is a novel proteasome interactor in mycobacteria and related organisms. eLife 7, e34055.10.7554/eLife.34055.028Search in Google Scholar
Zuhl, F., Seemuller, E., Golbik, R., and Baumeister, W. (1997). Dissecting the assembly pathway of the 20S proteasome. FEBS Lett. 418, 189–194.10.1016/S0014-5793(97)01370-7Search in Google Scholar
Zwickl, P., Ng, D., Woo, K.M., Klenk, H.P., and Goldberg, A.L. (1999). An archaebacterial ATPase, homologous to ATPases in the eukaryotic 26 S proteasome, activates protein breakdown by 20 S proteasomes. J. Biol. Chem. 274, 26008–26014.10.1074/jbc.274.37.26008Search in Google Scholar PubMed
©2020 Walter de Gruyter GmbH, Berlin/Boston
Articles in the same Issue
- Frontmatter
- Editorial
- Happy Birthday: Biological Chemistry is celebrating its 400th volume
- Reviews
- Bioenergetics and translational metabolism: implications for genetics, physiology and precision medicine
- Emerging mechanisms of drug-induced phospholipidosis
- Ribosome recycling in mRNA translation, quality control, and homeostasis
- Structure, dynamics and interactions of large SRP variants
- Space of Disse: a stem cell niche in the liver
- Structural investigations of cell-free expressed G protein-coupled receptors
- Coupling of import and assembly pathways in mitochondrial protein biogenesis
- Converting GTP hydrolysis into motion: versatile translational elongation factor G
- The Ras switch in structural and historical perspective
- Synthetic and biological approaches to map substrate specificities of proteases
- Proteasomes: unfoldase-assisted protein degradation machines
- Inorganic nitrite bioactivation and role in physiological signaling and therapeutics
Articles in the same Issue
- Frontmatter
- Editorial
- Happy Birthday: Biological Chemistry is celebrating its 400th volume
- Reviews
- Bioenergetics and translational metabolism: implications for genetics, physiology and precision medicine
- Emerging mechanisms of drug-induced phospholipidosis
- Ribosome recycling in mRNA translation, quality control, and homeostasis
- Structure, dynamics and interactions of large SRP variants
- Space of Disse: a stem cell niche in the liver
- Structural investigations of cell-free expressed G protein-coupled receptors
- Coupling of import and assembly pathways in mitochondrial protein biogenesis
- Converting GTP hydrolysis into motion: versatile translational elongation factor G
- The Ras switch in structural and historical perspective
- Synthetic and biological approaches to map substrate specificities of proteases
- Proteasomes: unfoldase-assisted protein degradation machines
- Inorganic nitrite bioactivation and role in physiological signaling and therapeutics

