Abstract
Objectives
Diabetic retinopathy (DR) is a common complication of diabetes mellitus (DM), with a risk of blindness. The study was to search for effective biomarkers of DR.
Methods
The key molecules of DR were mined by bioinformatics. Dual luciferase reporter gene assay, knockdown and overexpression assay were used to detect the regulatory relationship between hsa_circ_0053004, hsa-miR-646, and CBX2, and their clinical value was evaluated by constructing ROC curve. Levels of hsa_circ_0053004, hsa-miR-646, and CBX2 mRNA was detected by RT-qPCR, and western blotting was for detection of CBX2 protein expression. CCK-8 reagent and transwell assay were used for cell viability and migration detection.
Results
Hsa_circ_0053004 and CBX2 expressed highly in DR patients, while hsa-miR-646 was defective. Hsa_circ_0053004 down-regulated hsa-miR-646 by targeting it directly. Hsa-miR-646 inhibited CBX2 expression by targeting its 3′UTR. Overexpression of hsa_circ_0053004 or CBX2 promoted the high glucose-induced enhancement in human retinal microvascular endothelial cell (HRMEC) viability and migration, which was reversed by co-transfection of hsa-miR-646 mimic. The AUC of hsa_circ_0053004, hsa-miR-646, and CBX2 in predicting DR was 0.827, 0.805, and 0.817, respectively, and their combined AUC was 0.942.
Conclusions
Hsa_circ_0053004 promoted CBX2 expression by adsorbing hsa-miR-646, thereby promoting HRMECs to grow and migrate, and they may be good biomarkers of DR.
Introduction
The long-term increase in blood glucose caused by diabetes mellitus (DM) can lead to retinal vasculopathy, called diabetic retinopathy (DR) [1]. DR can be divided into non-proliferative DR (NDR) and proliferative DR (PDR) based on whether there is abnormal neovascularization emanating from the retina. The symptoms of NDR are relatively mild, with small bleeding spots, blood spots, hard oozing, and possible microangioma. Treatment at this stage is mainly to control blood glucose, oral administration of some drugs to improve microcirculation or hemostasis, and laser treatment can be used for macular edema [2], [3], [4]. PDR is the severe stage of DR, accompanied by the formation of new blood vessels. For the treatment of PDR, laser therapy can be applied to the initial stage to inhibit the growth of new blood vessels. If the laser treatment is not timely, it will lead to vitreous hemorrhage, the formation of the mechanical proliferative film, and retinal detachment, then the vitrectomy should be performed [5], [6], [7]. However, when there is too much proliferative membrane, the effect of surgery is not ideal. If DR is not treated in time, it will lead to blindness. Therefore, understanding the molecular mechanism of DR pathogenesis and seeking its valuable biomarkers will be very meaningful for the screening and intervention of DR.
Circular RNAs (circRNAs) are a class of circular non-coding RNA molecules. CircRNA is rich in the binding sites of microRNA (miRNA) and acts as a miRNA sponge in cells, thereby relieving the inhibition of miRNA on its target genes and increasing the expression level of target genes, which is called the competitive endogenous RNA (ceRNA) mechanism [8], 9]. MiRNAs are a class of short (approximately 22 nucleotides) non-coding single-stranded RNA molecules that are involved in post-transcriptional expression regulation of genes by destroying the stability of mRNA [10], 11]. The ceRNA mechanism in which circRNA participates is a vital regulatory mechanism of DR. CircMKLN1 has been reported to be abnormally high in the serum of patients with DR, and its inhibition was able to improve leakage of the retinal acellular capillary (RAC) in the DR mouse model. CircMKLN1 can also act as a molecular sponge to adsorb miR-26a-5p and regulate the expression of Rab11a, thereby mediating the autophagy induced by high glucose (HG)/methylglyoxal (MG) in human retinal microvascular endothelial cells (HRMECs) [12]. CircRNA_0084043 increases TGFA expression in retinal pigment epithelial cells (ARPE-19) through sponging miR-140-3p, which promotes HG-induced ARPE-19 damage [2]. Overexpression of circDMNT3B can reduce the RAC number of diabetic rats. Inhibition of circDMNT3B can hinder the expression of BAMBI by up-regulating miR-20b-5p, thus promoting the tube formation of HRMECs [13].
Here, we screened the candidate molecular markers of DR, hsa_circ_0053004, hsa-miR-646, and homo sapiens chromobox homolog 2 (CBX2), through bioinformatics, and explored their roles in the pathogenesis of DR.
Materials and methods
Bioinformatics analysis
The datasets of GSE193974 and GSE185011 were from the GEO database (https://www.ncbi.nlm.nih.gov/geo/). The downstream genes of hsa_circ_0053004 were predicted by Circular RNA Interactom (https://circinteractome.nia.nih.gov/), miRDB (https://mirdb.org/custom.html) and circBank (http://www.circbank.cn/searchCirc.html) databases. The target genes of hsa-miR-646 were predicted by the TargetScan8.0 (https://www.targetscan.org/) and miRDB databases. The binding site of hsa_circ_0053004 and hsa-miR-646 was from the Circular RNA Interactom database. The binding sequence of hsa-miR-646 and CBX2 mRNA was predicted by the TargetScan 8.0 database. The gene ontology (GO) analysis was performed in a Weishengxin platform (https://www.bioinformatics.com.cn/).
Patients
All participants in this retrospective study were from Army Center of PLA, including 168 patients with DM, 110 patients with NDR, and 102 patients with PDR. The clinical characteristics of all subjects were collected in Supplementary Table 1. The informed consent forms signed by all the volunteers and the samples from them were in good condition. This study was approved by the Ethics Committee of Army Center of PLA (No.20190201) and performed in line with the principles of the Declaration of Helsinki.
Cells
HRMECs were from Pricella biological company (China) and were cultured with the dedicated complete medium from the same company, at a 37 °C incubator with 5 % CO2. The cells were cultured with 5.5 mM glucose (normal glucose group, NG) or 25 mM glucose (high glucose group, HG).
RT-qPCR
Total RNA was isolated from cells by TRIzol Reagent (Invitrogen, USA). qPCR was performed in an ABI 7,500 system using SYBR®Green Realtime PCR Master Mix (Toyobo Biologics, China). The relative expression of genes was calculated by the ΔΔCT method. Hsa_circ_0053004 and CBX2 were normalized using GAPDH, and hsa-miR-646 was normalized by U6.
Detection of cell viability
The Cell Counting Kit-8 (CCK-8) kit (Solarbio, China) was used to assess cell viability. The 10 % CCK-8 solution prepared with the medium was added to the cells and stood still for 3 h. The absorbance at 450 nm was detected by an enzymoleter. The cell viability was calculated according to the instructions.
Detection of cell migration
Transwell assay was used to evaluate cell migration. Cells, starved for 12 h, were diluted with FBS-free medium containing 0.1 % BSA to a solution with a concentration of 1 × 105. The Transwell chamber (Corning, USA) containing 200 μL cell fluid was incubated in a 24-well plate containing the complete medium for 12 h. The migrated cells were fixed and stained with 4 % paraformaldehyde and 0.1 % crystal violet, respectively. Finally, the migrated cells were counted.
Dual luciferase reporter gene assay
Wild-type (wt) and mutant (mut) dual luciferase reporter vectors of hsa_circ_0053004 or CBX2 were co-transfected into HRMECs with hsa-miR-646 mimic or negative control (NC) and incubated for 24 h. Luciferase activity was checked by a luciferase reporter gene kit (Zeye Biotechnology LTD., China).
Western blotting
The protein samples were extracted using cell lysate. The protein concentration was detected by the BCA kit (Solarbio). Proteins were isolated by SDS-PAGE. The proteins were then transferred to the PVDF membrane (Millipore, Germany). PVDF membrane was incubated in 5 % skim milk powder for 1 h at room temperature. Then it was placed in the primary antibody to oscillate overnight at 4 °C. PVDF membrane was cleaned with TBST buffer for three times, then placed in the second antibody to incubate for 2 h at room temperature. Finally, the protein signal was detected by a chemiluminescence apparatus.
Evaluation of the predictive value of genes for DR
SPSS software was used to construct the receiver operating characteristic (ROC) curve and perform binary logistic regression analysis to evaluate the predictive value of genes for DR.
Statistical analysis
All data were analyzed using GraphPad Prism software and expressed as mean ± SD. The Student’s t-test was applied to measure the difference between the two groups. One-way ANOVA was used for multiple groups. p<0.05 was to be statistically remarkable.
Results
Hsa_circ_0053004 promoted HG-induced enhancement of HRMEC function
There were 584 circRNAs differentially expressed in patients with DM or DR, identified by GEO2R analysis in the GSE193974 dataset, including 115 down-regulated and 469 up-regulated (Figure 1A). Hsa_circ_0053004 expressed with the most multiple change (p=0.003, log2 FC=3.23), so we were interested in it. Hsa_circ_0053004 was up-regulated in the blood of DR patients, and it represented the more serious retinopathy (Figure 1B). HG increased the hsa_circ_0053004 expression in HRMECs (Figure 1C). The small interfering RNA of hsa_circ_0053004 (si-circ_0053004) significantly decreased the expression of hsa_circ_0053004, while its plasmid (ov-circ_0053004) obviously increased its level (Figure 1D). The down-regulation of hsa_circ_0053004 hindered the enhancement of HRMEC function induced by HG, and its up-regulation aggravated the effects of HG (Figure 1E and F).
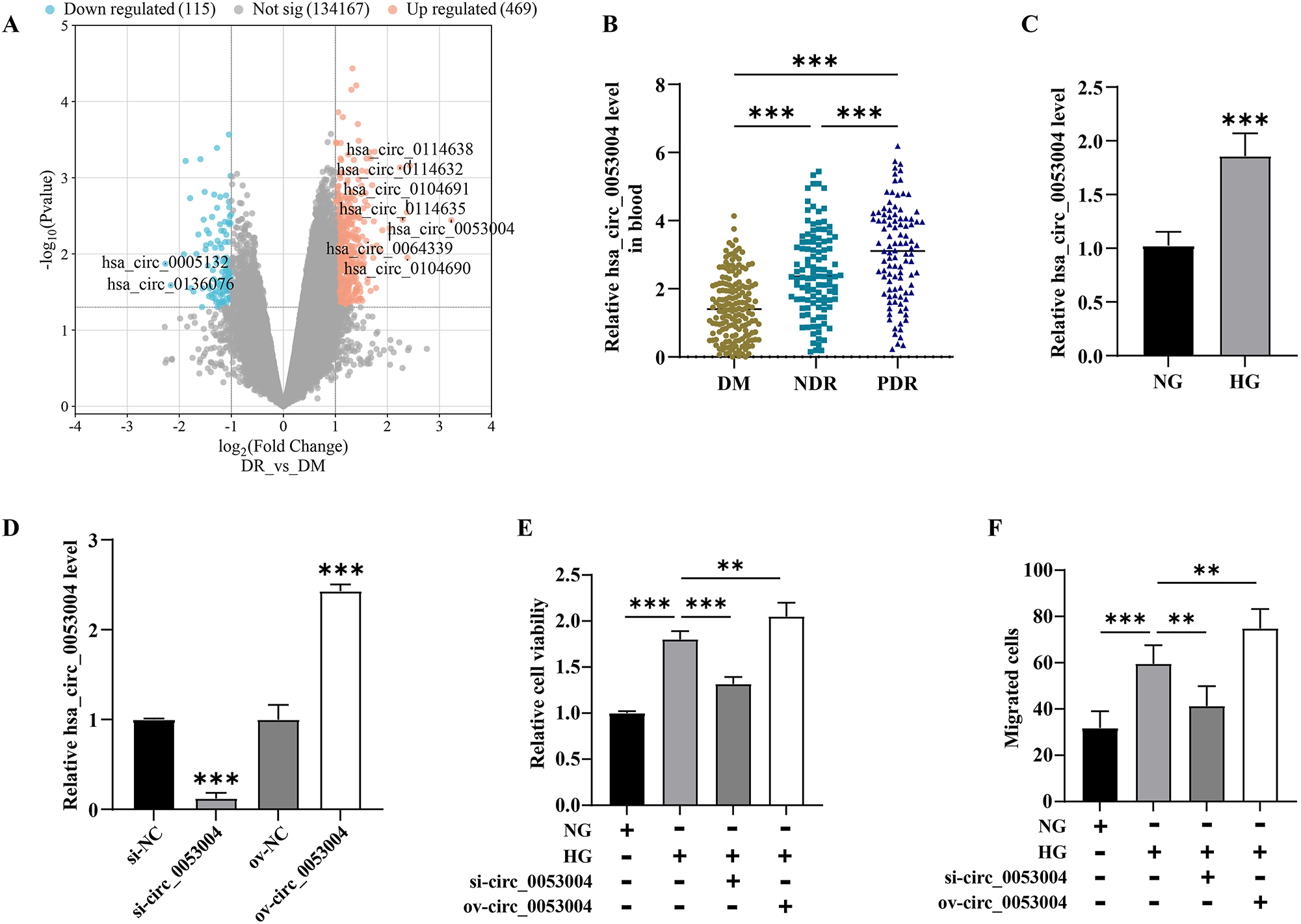
Hsa_circ_0053004 in HRMEC function. (A) Differentially expressed circRNAs between patients with DR and DM from the GSE19397 dataset. (B) Expression of hsa_circ_0053004 in the blood of patients. (C) Effect of HG on the expression of hsa_circ_0053004. (D) Efficiency of si-circ_0053004 and ov-circ_0053004. Effect of hsa_circ_0053004 on the cell viability (E) and migration (F) of HRMECs. **p<0.01; ***p<0.001.
Hsa-miR-646 eliminated the enhancement of hsa_circ_0053004 on HRMEC function
The downstream miRNAs of hsa_circ_0053004 were predicted by different databases, in which hsa-miR-646 was shared (Figure 2A). Bioinformatics analysis revealed the binding sequence of hsa_circ_0053004 and hsa-miR-646 (Supplementary Figure 1A). Hsa-miR-646 significantly reduced the luciferase activity of cells transfected with the wt-circ_0053004 luciferase vector; however, no significant changes were observed in cells transfected with mut-circ_0053004 (Figure 2B). Knockdown of hsa_circ_0053004 decreased the level of hsa-miR-646 in cells, and its overexpression increased the hsa-miR-646 level (Figure 2C).

Hsa_circ_0053004 promoted the function of HRMECs by regulating hsa-miR-646. (A) The downstream miRNAs of hsa_circ_0053004 predicted by different databases. (B) Effect of hsa-miR-646 mimic on the luciferase activity of HRMECs transfected with wt-circ_0053004 or mut-circ_0053004. (C) Hsa_circ_0053004 regulated the level of hsa-miR-646 in HRMECs. (D) Hsa-miR-646 level in the blood of DM and DR patients (E) Effect of HG on hsa-miR-646 expression. (F-G) Efficiency of hsa-miR-646 mimic and inhibitor. (H-I) Effect of hsa-miR-646 on the cell viability and migration of HRMECs. **p<0.01; ***p<0.001.
The expression of hsa-miR-646 was deficient in the blood of DR patients (Figure 2D). HG decreased the hsa-miR-646 expression of HRMECs (Figure 2E). Hsa-miR-646 mimic (mim-646) and inhibitor were introduced for functional study (Figure 2F and G). Overexpression of hsa-miR-646 reversed the increased ability to proliferate and migrate in HRMECs induced by HG, and inhibition of hsa-miR-646 contributed to HG-induced cell proliferation and migration. Moreover, the increase of hsa-miR-646 level reversed the enhanced function of HRMECs induced by ov-circ_0053004 transfection (Figure 2H and I), suggesting that hsa_circ_0053004 promoted HRMECs to grow and migrate by negatively regulating hsa-miR-646.
CBX2 was the target of hsa-miR-646
There were 9,088 and 880 downstream genes of hsa-miR-646 were predicted by the TargetScan 8.0 database and miRDB database, respectively, among which 27 genes were identified to be the differentially expressed gene in DR and DM patients through the GEO2R analysis in the GSE185011 dataset (Supplementary Figure 1B). The GO analysis results of these 27 genes showed that CBX2 was a component of the polycomb repressive complex 1 (PRC1) (Figure 3A). Other components of PRC1 have been reported to be associated with insulin sensitivity [14], so we were very interested in the role of CBX2 in DR. The gene expression profile in the GSE185011 dataset revealed that CBX2 expressed highly in patients with DR compared to DM patients (Figure 3B), indicating that CBX2 may be a risk factor for DR. The binding sequence of hsa-miR-646 and CBX2 mRNA was displayed in the TargetScan 8.0 database (Supplementary Figure 1C). The mimic of hsa-miR-646 reduced luciferase activity in cells transfected with wt-CBX2, but hardly affected cells transfected with mut-CBX2 (Figure 3C). Moreover, hsa-miR-646 mimic significantly inhibited CBX2 expression (Figure 3D and E).
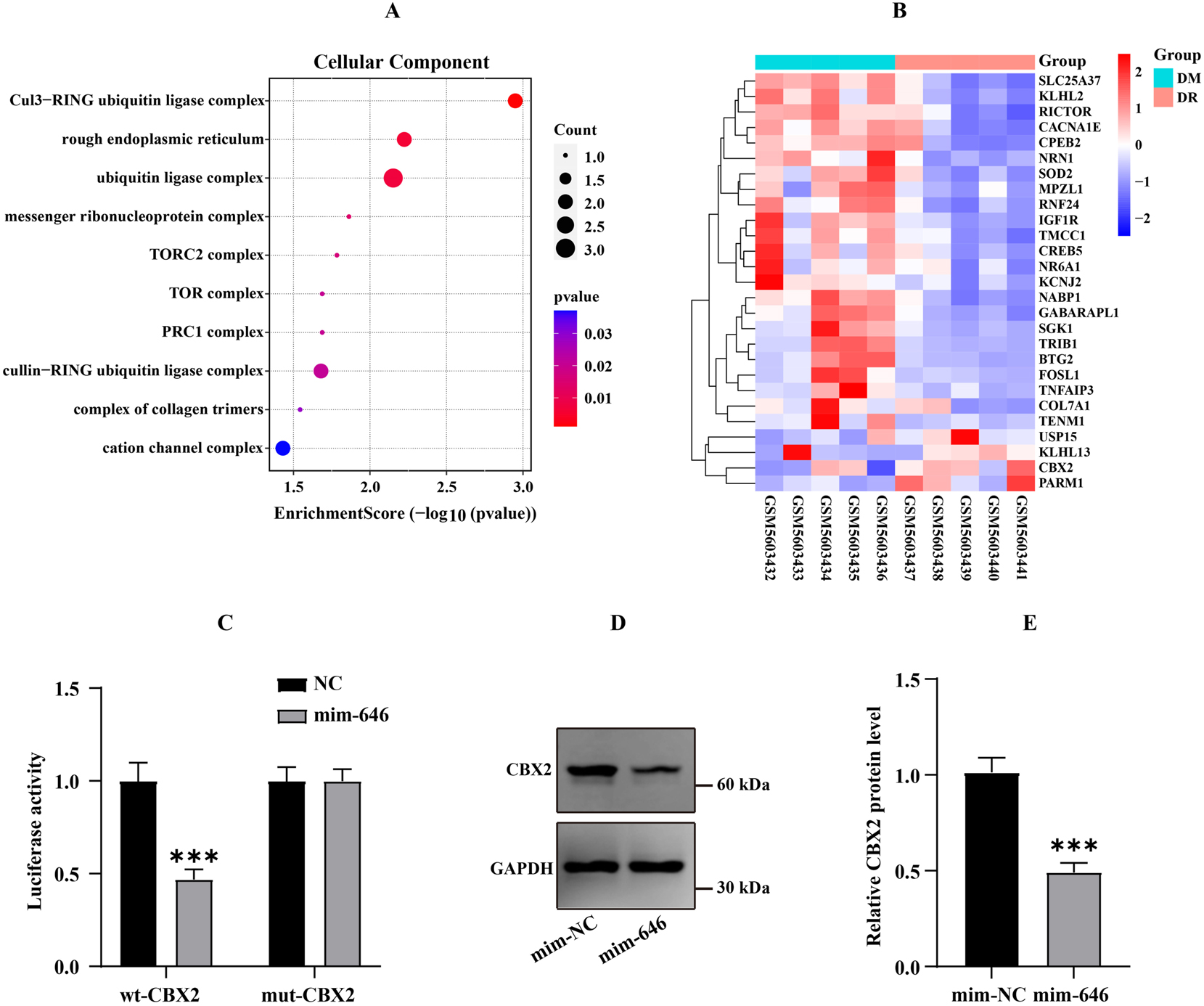
CBX2 was a target gene of hsa-miR-646 (A) The GO analysis of common genes. (B) The expression map of common genes in GSE185011 dataset. (C) Effect of hsa-miR-646 mimic on the luciferase activity of HRMECs transfected with wt-CBX2 or mut-CBX2 (D-E) Hsa-miR-646 regulated CBX2 expression. ***p<0.001.
CBX2 aggravated HG-induced enhanced function of HRMECs
The expression of CBX2 was up-regulated in the blood of DR patients (Figure 4A). HG led to increased expression of CBX2 (Figure 4B and C). Transfecting plasmid of CBX2 (ov-CBX2) markedly promoted its expression (Figure 4D and E), which aggravated the HG-induced enhancement in HRMEC proliferation and migration, which was reversed by overexpression of hsa-miR-646 (Figure 4F and G).

CBX2 aggravated HG-induced enhancement of HRMEC function. (A) CBX2 expression in the blood of DM and DR patients. (B-C) HG promoted the CBX2 expression. (D-E) The working efficiency of ov-CBX2. (F-G) Effects of the co-transfection of ov-CBX2 and hsa-miR-646 mimic on the HRMEC viability and migration. ***p<0.001.
Prediction function of hsa_circ_0053004/hsa-miR-646/CBX2 axis for DR
The ROC curve showed that hsa_circ_0053004, hsa-miR-646, and CBX2 were good indicators for predicting the occurrence of DR, with AUC of 0.872, 0.805, and 0.817, respectively (Figure 5A–C). The combination of three indicators showed a higher accuracy in predicting the occurrence of DR (AUC = 0.942) (Figure 5D). In addition, multivariate logistic regression analysis showed that hsa_circ_0053004, hsa-miR-646, and CBX2 were independent influencing factors of DR (Table 1). Among them, hsa_circ_0053004 and CBX2 were risk factors, while hsa-miR-646 was a protective factor for DR.

Prediction function of hsa_circ_0053004/hsa-miR-646/CBX2 axis for DR. The ROC curve of hsa_circ_0053004 (A), hsa-miR-646 (B), CBX2 (C), and their combination (D) in predicting DR. AUC, area under curve.
Multivariate logistic regression analysis.
| Group | OR | 95 % CI | p-Value | |
|---|---|---|---|---|
| Lower | Upper | |||
| Hsa_circ_0053004 | 2.870 | 2.072 | 3.977 | <0.001 |
| Hsa-miR-646 | 0.593 | 0.499 | 0.706 | <0.001 |
| CBX2 | 2.265 | 1.754 | 2.927 | <0.001 |
| Gender | 1.374 | 0.704 | 2.681 | 0.352 |
| Age | 1.476 | 0.760 | 2.866 | 0.250 |
| Smoking | 1.370 | 0.700 | 2.682 | 0.358 |
| Drinking | 1.067 | 0.547 | 2.081 | 0.849 |
| Obesity | 1.026 | 0.523 | 2.012 | 0.941 |
| Hypertension | 1.293 | 0.652 | 2.567 | 0.462 |
-
OR, odds ratio; CI, confidence interval.
Discussion
DR, a type of retinal microvascular disease, is the outcome of almost all patients with DM. In severe cases, DR will be due to retinal detachment, manifesting as blurred vision, floaters, distorted vision, and even partial or total vision loss [15], 16]. If DR patients can be treated timely, these conditions will be completely avoided. Therefore, exploring the molecular mechanism of DR development and mining its effective biomarkers can timely screen DR for early intervention and prevent the occurrence of severe DR.
The circRNA-mediated ceRNA mechanism is an important participant in the pathogenesis of DR [2], 12], 13]. In this study, we screened the key molecules of DR, hsa_circ_0053004, hsa-miR-646, and CBX2, through bioinformatics. Compared with DM patients without DR, hsa_circ_0053004, and CBX2 were abnormally high-expressed in the peripheral blood of DR patients, while hsa-miR-646 expression was scarce. Bioinformatics analysis showed that hsa_circ_0053004 had the binding site of hsa-miR-646, which was consistent with the site of hsa-miR-646 targeting CBX2 mRNA. Hsa_circ_0053004 bound to hsa-miR-646 by competing with CBX2 mRNA, promoting the expression of CBX2. Overexpression of hsa_circ_0053004 aggravated HG-induced enhancement of HRMEC function by regulating the expression of CBX2 through adsorption of hsa-miR-646. The ROC curve and multivariate logistic regression analysis showed that hsa_circ_0053004 and CBX2 were risk factors for the development of DM into DR, while hsa-miR-646 was a protective factor. The combination of them can predict the occurrence of DR accurately (AUC = 0.942).
Studies have shown that miRNAs are potential biomarkers for DR. The expression profiles of exosomal miRNA from the serum of DM or DR patients analyzed by RNA sequencing revealed 18 up-regulated miRNAs in DR. The overexpression of miR-3976 promoted the apoptosis of mouse retinal ganglion cell 5 (RGC-5) [17]. The level of miR-23a was significantly reduced in blood and tear samples from DR patients compared to controls. Overexpression of miR-23a can inhibit the proliferative activity of HRMECs by decreasing the expression level of vascular endothelial growth factor (VEGF) [18]. Studies have shown that the knockdown of circCOL1A2 can inhibit HG-induced migration, proliferation, and angiogenesis of HRMECs by up-regulating hsa-miR-646 and inhibiting fibroblast growth factor 7 (FGF7) [19]. In addition, hsa_circ_0041795 can activate VEGFC by sponging hsa-miR-646, enhancing the damage of ARPE-19 viability induced by HG [20]. We demonstrated that hsa-miR-646 was low-expressed in the blood of DR patients, and its overexpression reversed the HG-induced enhancement of the migration and cell viability in HRMECs. Hsa_circ_0053004 promoted the HG-induced heightener of HRMEC function by up-regulating the expression of CBX2 through sponging hsa-miR-646.
Our GO analysis displayed that CBX2 was a component of PRC1. Bmi1, another component of PRC1, has been reported to play a crucial role in maintaining glucose homeostasis, with an inverse correlation between its level and insulin sensitivity [14], suggesting that PRC1 may be the key pathogenic factor of DM. Our experimental results show that CBX2 was up-regulated in DR patients, and its overexpression significantly promoted the HG-induced enhancement of HRMEC function.
Conclusions
In conclusion, we revealed the role of the hsa_circ_0053004/hsa-miR-646/CBX2 axis in HRMEC function, which may be an important mechanism for the development of DR. Our study will provide theoretical support for the screening and intervention of DR.
-
Research ethics: This study was approved by the Ethics Committee of Army Center of PLA (No.20190201) and performed in line with the principles of the Declaration of Helsinki.
-
Informed consent: The informed consent forms signed by all the volunteers and the samples from them were in good condition.
-
Author contributions: All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by S. Z., M. L. and Y. C. The first draft of the manuscript was written by S. Z. and M. L. and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
-
Use of Large Language Models, AI and Machine Learning Tools: None.
-
Conflict of interest: There is no conflict of interest in this study.
-
Research funding: This research received no specific grant from any funding agency in the public, commercial or not-for-profit sectors.
-
Data availability: The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
References
1. Tan, TE, Wong, TY. Diabetic retinopathy: looking forward to 2030. Front Endocrinol (Lausanne) 2022;13:1077669. https://doi.org/10.3389/fendo.2022.1077669.Search in Google Scholar PubMed PubMed Central
2. Lin, KY, Hsih, WH, Lin, YB, Wen, CY, Chang, TJ. Update in the epidemiology, risk factors, screening, and treatment of diabetic retinopathy. J Diabetes Investig 2021;12:1322–5. https://doi.org/10.1111/jdi.13480.Search in Google Scholar PubMed PubMed Central
3. Honasoge, A, Nudleman, E, Smith, M, Rajagopal, R. Emerging insights and interventions for diabetic retinopathy. Curr Diab Rep 2019;19:100. https://doi.org/10.1007/s11892-019-1218-2.Search in Google Scholar PubMed PubMed Central
4. Mishra, S, Vishwakarma, PK, Tripathi, M, Ojha, S, Tripathi, SM. Diabetic retinopathy: clinical features, risk factors, and treatment options. Curr Diabetes Rev 2024;20. https://doi.org/10.2174/0115733998252551231018080419. e271023222871.Search in Google Scholar PubMed
5. Chaudhary, S, Zaveri, J, Becker, N. Proliferative diabetic retinopathy (PDR). Dis Mon 2021;67:101140. https://doi.org/10.1016/j.disamonth.2021.101140.Search in Google Scholar PubMed
6. Chen, SN, Chen, SJ, Wu, TT, Wu, WC, Yang, CH, Yang, CM. Refining vitrectomy for proliferative diabetic retinopathy. Graefes Arch Clin Exp Ophthalmol 2023;261:3659–70. https://doi.org/10.1007/s00417-023-06134-w.Search in Google Scholar PubMed PubMed Central
7. Studnicka, J, Nemcansky, J, Vyslouzilova, D, Ernest, J, Nemec, P. Diabetic retinopathy – diagnostics and treatment guidelines. Ceska a Slov Oftalmol 2023;79:238–47. https://doi.org/10.31348/2023/28.Search in Google Scholar PubMed
8. Tay, Y, Rinn, J, Pandolfi, PP. The multilayered complexity of ceRNA crosstalk and competition. Nature 2014;505:344–52. https://doi.org/10.1038/nature12986.Search in Google Scholar PubMed PubMed Central
9. Zhou, WY, Cai, ZR, Liu, J, Wang, DS, Ju, HQ, Xu, RH. Circular RNA: metabolism, functions and interactions with proteins. Mol Cancer 2020;19:172. https://doi.org/10.1186/s12943-020-01286-3.Search in Google Scholar PubMed PubMed Central
10. Pala, M. Exercise and microrna. Georgian Med News 2023:146–53.Search in Google Scholar
11. Hwang, H, Chang, HR, Baek, D. Determinants of functional MicroRNA targeting. Mol Cells 2023;46:21–32. https://doi.org/10.14348/molcells.2023.2157.Search in Google Scholar PubMed PubMed Central
12. Yang, J, Tan, C, Wang, Y, Zong, T, Xie, T, Yang, Q, et al.. The circRNA MKLN1 regulates autophagy in the development of diabetic retinopathy. Biochim Biophys Acta, Mol Basis Dis 2023;1869:166839. https://doi.org/10.1016/j.bbadis.2023.166839.Search in Google Scholar PubMed
13. Zhu, K, Hu, X, Chen, H, Li, F, Yin, N, Liu, AL, et al.. Downregulation of circRNA DMNT3B contributes to diabetic retinal vascular dysfunction through targeting miR-20b-5p and BAMBI. EBioMedicine 2019;49:341–53. https://doi.org/10.1016/j.ebiom.2019.10.004.Search in Google Scholar PubMed PubMed Central
14. Cannon, CE, Titchenell, PM, Groff, DN, El Ouaamari, A, Kulkarni, RN, Birnbaum, MJ, et al.. The Polycomb protein, Bmi1, regulates insulin sensitivity. Mol Metabol 2014;3:794–802. https://doi.org/10.1016/j.molmet.2014.08.002.Search in Google Scholar PubMed PubMed Central
15. Bryl, A, Mrugacz, M, Falkowski, M, Zorena, K. The effect of diet and lifestyle on the course of diabetic retinopathy-A review of the literature. Nutrients 2022;14. https://doi.org/10.3390/nu14061252.Search in Google Scholar PubMed PubMed Central
16. Do, DV, Han, G, Abariga, SA, Sleilati, G, Vedula, SS, Hawkins, BS. Blood pressure control for diabetic retinopathy. Cochrane Database Syst Rev 2023;3:CD006127. https://doi.org/10.1002/14651858.CD006127.pub3.Search in Google Scholar PubMed PubMed Central
17. Yang, S, Zhang, J, Zeng, T, Zheng, J, Min, J, Chen, L. Role of circulating exosomal miRNA-3976 in early diabetic retinopathy. Int J Nanomed 2023;18:3695–709. https://doi.org/10.2147/IJN.S414393.Search in Google Scholar PubMed PubMed Central
18. Sun, L, Liu, X, Zuo, Z. Regulatory role of miRNA-23a in diabetic retinopathy. Exp Ther Med 2021;22:1477. https://doi.org/10.3892/etm.2021.10912.Search in Google Scholar PubMed PubMed Central
19. Cao, H, Xu, X, Wang, K, Li, C. Down-regulation of circCOL1A2 suppresses the dysfunction of diabetes-related retinal microvascular endothelial cells via miR-646/FGF7 Axis. Curr Eye Res 2022;47:1525–33. https://doi.org/10.1080/02713683.2022.2110264.Search in Google Scholar PubMed
20. Sun, H, Kang, X. Hsa_circ_0041795 contributes to human retinal pigment epithelial cells (ARPE 19) injury induced by high glucose via sponging miR-646 and activating VEGFC. Gene 2020;747:144654. https://doi.org/10.1016/j.gene.2020.144654.Search in Google Scholar PubMed
Supplementary Material
This article contains supplementary material (https://doi.org/10.1515/tjb-2024-0329).
© 2025 the author(s), published by De Gruyter, Berlin/Boston
This work is licensed under the Creative Commons Attribution 4.0 International License.
Articles in the same Issue
- Frontmatter
- Review Article
- Unveiling the hidden clinical and economic impact of preanalytical errors
- Research Articles
- To explore the role of hsa_circ_0053004/hsa-miR-646/CBX2 in diabetic retinopathy based on bioinformatics analysis and experimental verification
- Study on the LINC00578/miR-495-3p/RNF8 axis regulating breast cancer progression
- Comparison of two different anti-mullerian hormone measurement methods and evaluation of anti-mullerian hormone in polycystic ovary syndrome
- The evaluation of the relationship between anti angiotensin type I antibodies in hypertensive patients undergoing kidney transplantation
- Evaluation of neopterin, oxidative stress, and immune system in silicosis
- Assessment of lipocalin-1, resistin, cathepsin-D, neurokinin A, agmatine, NGF, and BDNF serum levels in children with Autism Spectrum Disorder
- Regulatory nexus in inflammation, tissue repair and immune modulation in Crimean-Congo hemorrhagic fever: PTX3, FGF2 and TNFAIP6
- Pasteur effect in leukocyte energy metabolism of patients with mild, moderate, and severe COVID-19
- Thiol-disulfide homeostasis and ischemia-modified albumin in patients with sepsis
- Myotonic dystrophy type 1 and oxidative imbalance: evaluation of ischemia-modified albumin and oxidant stress
- Antioxidant and alpha-glucosidase inhibitory activities of flavonoids isolated from fermented leaves of Camellia chrysantha (Hu) Tuyama
- Examination of the apelin signaling pathway in acetaminophen-induced hepatotoxicity in rats
- Integrating network pharmacology, in silico molecular docking and experimental validation to explain the anticancer, apoptotic, and anti-metastatic effects of cosmosiin natural product against human lung carcinoma
- Validation of Protein A chromatography: orthogonal method with size exclusion chromatography validation for mAb titer analysis
- The evaluation of the efficiency of Atellica UAS800 in detecting pathogens (rod, cocci) causing urinary tract infection
- Case Report
- Exploring inherited vitamin B responsive disorders in the Moroccan population: cutting-edge diagnosis via GC-MS profiling
- Letter to the Editor
- Letter to the Editor: “Gene mining, recombinant expression and enzymatic characterization of N-acetylglucosamine deacetylase”
Articles in the same Issue
- Frontmatter
- Review Article
- Unveiling the hidden clinical and economic impact of preanalytical errors
- Research Articles
- To explore the role of hsa_circ_0053004/hsa-miR-646/CBX2 in diabetic retinopathy based on bioinformatics analysis and experimental verification
- Study on the LINC00578/miR-495-3p/RNF8 axis regulating breast cancer progression
- Comparison of two different anti-mullerian hormone measurement methods and evaluation of anti-mullerian hormone in polycystic ovary syndrome
- The evaluation of the relationship between anti angiotensin type I antibodies in hypertensive patients undergoing kidney transplantation
- Evaluation of neopterin, oxidative stress, and immune system in silicosis
- Assessment of lipocalin-1, resistin, cathepsin-D, neurokinin A, agmatine, NGF, and BDNF serum levels in children with Autism Spectrum Disorder
- Regulatory nexus in inflammation, tissue repair and immune modulation in Crimean-Congo hemorrhagic fever: PTX3, FGF2 and TNFAIP6
- Pasteur effect in leukocyte energy metabolism of patients with mild, moderate, and severe COVID-19
- Thiol-disulfide homeostasis and ischemia-modified albumin in patients with sepsis
- Myotonic dystrophy type 1 and oxidative imbalance: evaluation of ischemia-modified albumin and oxidant stress
- Antioxidant and alpha-glucosidase inhibitory activities of flavonoids isolated from fermented leaves of Camellia chrysantha (Hu) Tuyama
- Examination of the apelin signaling pathway in acetaminophen-induced hepatotoxicity in rats
- Integrating network pharmacology, in silico molecular docking and experimental validation to explain the anticancer, apoptotic, and anti-metastatic effects of cosmosiin natural product against human lung carcinoma
- Validation of Protein A chromatography: orthogonal method with size exclusion chromatography validation for mAb titer analysis
- The evaluation of the efficiency of Atellica UAS800 in detecting pathogens (rod, cocci) causing urinary tract infection
- Case Report
- Exploring inherited vitamin B responsive disorders in the Moroccan population: cutting-edge diagnosis via GC-MS profiling
- Letter to the Editor
- Letter to the Editor: “Gene mining, recombinant expression and enzymatic characterization of N-acetylglucosamine deacetylase”


